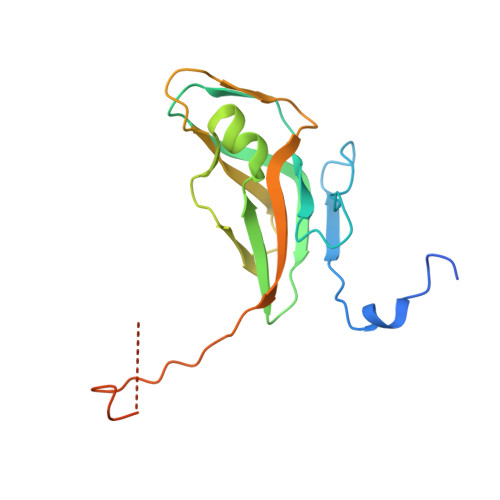
Direct contacts between conserved motifs of different subunits provide major contribution to active site organization in human and mycobacterial dUTPases.
Takacs, E., Nagy, G., Leveles, I., Harmat, V., Lopata, A., Toth, J., Vertessy, B.G.(2010) FEBS Lett 584: 3047-3054
- PubMed: 20493855 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.febslet.2010.05.018
- Primary Citation Related Structures:
3H6D, 3I93 - PubMed Abstract:
dUTP pyrophosphatases (dUTPases) are essential for genome integrity. Recent results allowed characterization of the role of conserved residues. Here we analyzed the Asp/Asn mutation within conserved Motif I of human and mycobacterial dUTPases, wherein the Asp residue was previously implicated in Mg(2+)-coordination. Our results on transient/steady-state kinetics, ligand binding and a 1.80 A resolution structure of the mutant mycobacterial enzyme, in comparison with wild type and C-terminally truncated structures, argue that this residue has a major role in providing intra- and intersubunit contacts, but is not essential for Mg(2+) accommodation. We conclude that in addition to the role of conserved motifs in substrate accommodation, direct subunit interaction between protein atoms of active site residues from different conserved motifs are crucial for enzyme function.
- Institute of Enzymology, BRC, Hungarian Academy of Sciences, Budapest, Hungary.
Organizational Affiliation: