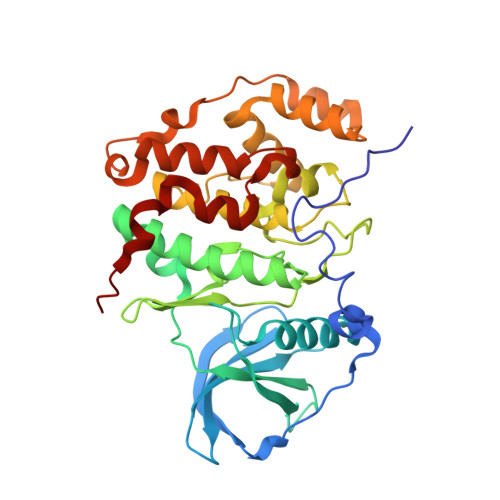
The CK2alpha/CK2beta Interface of Human Protein Kinase CK2 Harbors a Binding Pocket for Small Molecules
Raaf, J., Brunstein, E., Issinger, O.-G., Niefind, K.(2008) Chem Biol 15: 111-117
- PubMed: 18291315 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2007.12.012
- Primary Citation Related Structures:
3H30 - PubMed Abstract:
The Ser/Thr kinase CK2 (previously called casein kinase 2) is composed of two catalytic chains (CK2 alpha) attached to a dimer of noncatalytic subunits (CK2 beta). CK2 is involved in suppression of apoptosis, cell survival, and tumorigenesis. To investigate these activities and possibly affect them, selective CK2 inhibitors are required. An often-used CK2 inhibitor is 5,6-dichloro-1-beta-D-ribofuranosylbenzimidazole (DRB). In a complex structure with human CK2 alpha, DRB binds to the canonical ATP cleft, but additionally it occupies an allosteric site that can be alternatively filled by glycerol. Inhibition kinetic studies corroborate the dual binding mode of the inhibitor. Structural comparisons reveal a surprising conformational plasticity of human CK2 alpha around both DRB binding sites. After local rearrangement, the allosteric site serves as a CK2 beta interface. This opens the potential to construct molecules interfering with the CK2 alpha/CK2 beta interaction.
- Institut für Biochemie, Universität zu Köln, Zülpicher Str. 47, D-50674 Köln, Germany.
Organizational Affiliation: