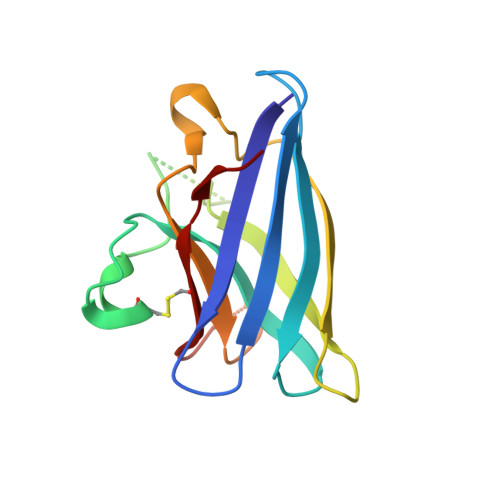
Structures of Pathogenic SOD1 Mutants H80R and D124V: Disrupted Zinc-binding and Compromised Post-translational Modification by the Copper Chaperone CCS
Seetharaman, S.V., Winkler, D.D., Taylor, A.B., Cao, X., Whitson, L.J., Doucette, P.A., Valentine, J.S., Carroll, M.C., Culotta, V.C., Hart, P.J.To be published.