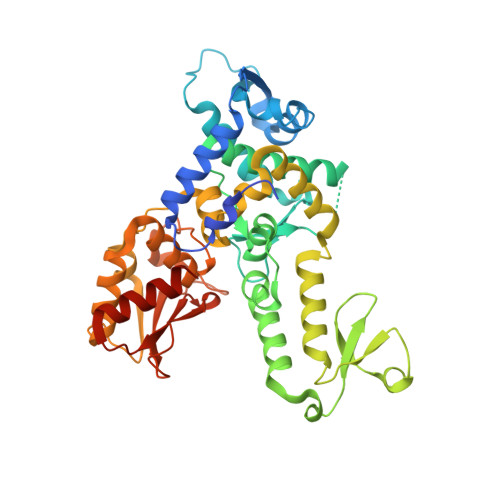
A structural element within the HUWE1 HECT domain modulates self-ubiquitination and substrate ubiquitination activities.
Pandya, R.K., Partridge, J.R., Love, K.R., Schwartz, T.U., Ploegh, H.L.(2010) J Biological Chem 285: 5664-5673
- PubMed: 20007713 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.051805
- Primary Citation Related Structures:
3H1D - PubMed Abstract:
E3 ubiquitin ligases catalyze the final step of ubiquitin conjugation and regulate numerous cellular processes. The HECT class of E3 ubiquitin (Ub) ligases directly transfers Ub from bound E2 enzyme to a myriad of substrates. The catalytic domain of HECT Ub ligases has a bilobal architecture that separates the E2 binding region and catalytic site. An important question regarding HECT domain function is the control of ligase activity and specificity. Here we present a functional analysis of the HECT domain of the E3 ligase HUWE1 based on crystal structures and show that a single N-terminal helix significantly stabilizes the HECT domain. We observe that this element modulates HECT domain activity, as measured by self-ubiquitination induced in the absence of this helix, as distinct from its effects on Ub conjugation of substrate Mcl-1. Such subtle changes to the protein may be at the heart of the vast spectrum of substrate specificities displayed by HECT domain E3 ligases.
- Whitehead Institute for Biomedical Research, Cambridge, Massachusetts 02142, USA.
Organizational Affiliation: