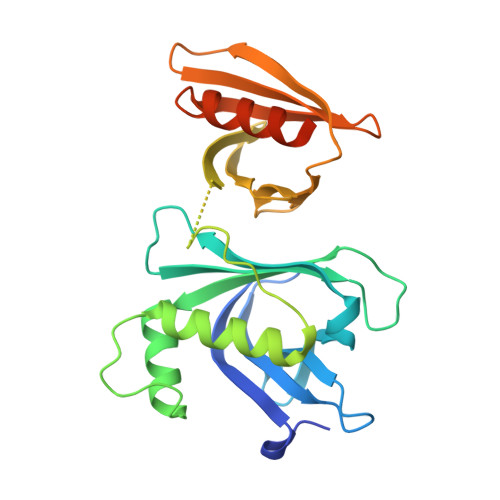
Structural analysis of Rtt106p reveals a DNA-binding role required for heterochromatin silencing
Liu, Y., Huang, H., Zhou, B.O., Wang, S.S., Hu, Y., Li, X., Liu, J., Zang, J., Niu, L., Wu, J., Zhou, J.Q., Teng, M., Shi, Y.(2010) J Biological Chem 285: 4251-4262
- PubMed: 20007951 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.055996
- Primary Citation Related Structures:
3GYO, 3GYP - PubMed Abstract:
Rtt106p is a Saccharomyces cerevisiae histone chaperone with roles in heterochromatin silencing and nucleosome assembly. The molecular mechanism by which Rtt106p engages in chromatin dynamics remains unclear. Here, we report the 2.5 A crystal structure of the core domain of Rtt106p, which adopts an unusual "double pleckstrin homology" domain architecture that represents a novel structural mode for histone chaperones. A histone H3-H4-binding region and a novel double-stranded DNA-binding region have been identified. Mutagenesis studies reveal that the histone and DNA binding activities of Rtt106p are involved in Sir protein-mediated heterochromatin formation. Our results uncover the structural basis of the diverse functions of Rtt106p and provide new insights into its cellular roles.
- From the Hefei National Laboratory for Physical Sciences at Microscale and School of Life Science, University of Science and Technology of China, Hefei, Anhui 230026 and.
Organizational Affiliation: