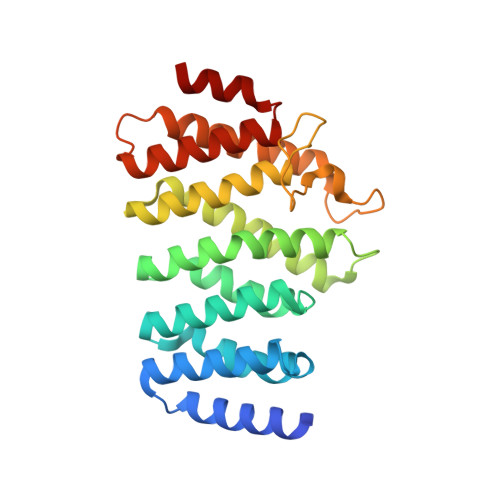
Crystal structure of the HEAT domain from the Pre-mRNA processing factor Symplekin.
Kennedy, S.A., Frazier, M.L., Steiniger, M., Mast, A.M., Marzluff, W.F., Redinbo, M.R.(2009) J Mol Biology 392: 115-128
- PubMed: 19576221
- DOI: https://doi.org/10.1016/j.jmb.2009.06.062
- Primary Citation Related Structures:
3GS3 - PubMed Abstract:
The majority of eukaryotic pre-mRNAs are processed by 3'-end cleavage and polyadenylation, although in metazoa the replication-dependent histone mRNAs are processed by 3'-end cleavage but not polyadenylation. The macromolecular complex responsible for processing both canonical and histone pre-mRNAs contains the approximately 1160-residue protein Symplekin. Secondary-structural prediction algorithms identified putative HEAT domains in the 300 N-terminal residues of all Symplekins of known sequence. The structure and dynamics of this domain were investigated to begin elucidating the role Symplekin plays in mRNA maturation. The crystal structure of the Drosophila melanogaster Symplekin HEAT domain was determined to 2.4 A resolution with single-wavelength anomalous dispersion phasing methods. The structure exhibits five canonical HEAT repeats along with an extended 31-amino-acid loop (loop 8) between the fourth and fifth repeat that is conserved within closely related Symplekin sequences. Molecular dynamics simulations of this domain show that the presence of loop 8 dampens correlated and anticorrelated motion in the HEAT domain, therefore providing a neutral surface for potential protein-protein interactions. HEAT domains are often employed for such macromolecular contacts. The Symplekin HEAT region not only structurally aligns with several established scaffolding proteins, but also has been reported to contact proteins essential for regulating 3'-end processing. Together, these data support the conclusion that the Symplekin HEAT domain serves as a scaffold for protein-protein interactions essential to the mRNA maturation process.
- Department of Chemistry, University of North Carolina at Chapel Hill, 27599, USA.
Organizational Affiliation: