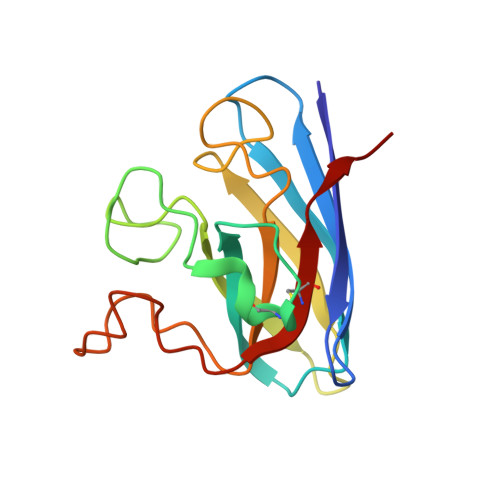
Structural and biophysical properties of the pathogenic SOD1 variant H46R/H48Q.
Winkler, D.D., Schuermann, J.P., Cao, X., Holloway, S.P., Borchelt, D.R., Carroll, M.C., Proescher, J.B., Culotta, V.C., Hart, P.J.(2009) Biochemistry 48: 3436-3447
- PubMed: 19227972 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi8021735
- Primary Citation Related Structures:
3GQF - PubMed Abstract:
Over 100 mutations in the gene encoding human copper-zinc superoxide dismutase (SOD1) cause an inherited form of the fatal neurodegenerative disease amyotrophic lateral sclerosis (ALS). Two pathogenic SOD1 mutations, His46Arg (H46R) and His48Gln (H48Q), affect residues that act as copper ligands in the wild type enzyme. Transgenic mice expressing a human SOD1 variant containing both mutations develop paralytic disease akin to ALS. Here we show that H46R/H48Q SOD1 possesses multiple characteristics that distinguish it from the wild type. These properties include the following: (1) an ablated copper-binding site, (2) a substantially weakened affinity for zinc, (3) a binding site for a calcium ion, (4) the ability to form stable heterocomplexes with the copper chaperone for SOD1 (CCS), and (5) compromised CCS-mediated oxidation of the intrasubunit disulfide bond in vivo. The results presented here, together with data on pathogenic SOD1 proteins coming from cell culture and transgenic mice, suggest that incomplete posttranslational modification of nascent SOD1 polypeptides via CCS may be a characteristic shared by familial ALS SOD1 mutants, leading to a population of destabilized, off-pathway folding intermediates that are toxic to motor neurons.
- Department of Biochemistry and X-ray Crystallography Core Laboratory, the University of Texas Health Science Center, San Antonio, Texas 78229-3900, USA.
Organizational Affiliation: