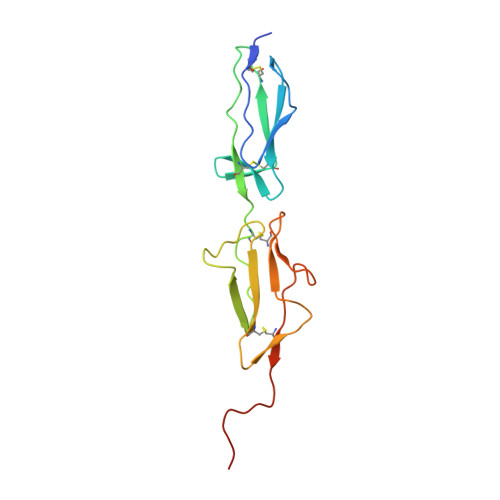
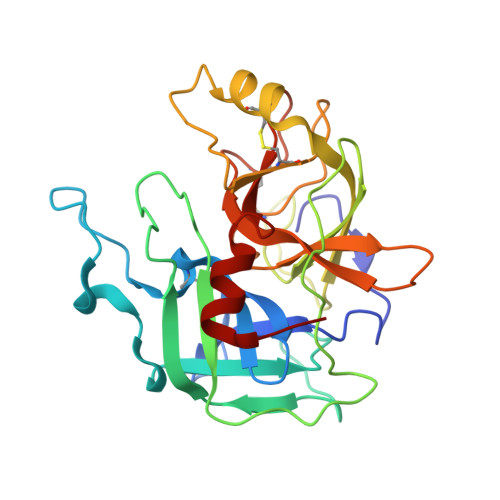
MASP-1, a promiscuous complement protease: structure of its catalytic region reveals the basis of its broad specificity.
Dobo, J., Harmat, V., Beinrohr, L., Sebestyen, E., Zavodszky, P., Gal, P.(2009) J Immunol 183: 1207-1214
- PubMed: 19564340 Search on PubMed
- DOI: https://doi.org/10.4049/jimmunol.0901141
- Primary Citation Related Structures:
3GOV - PubMed Abstract:
Mannose-binding lectin (MBL)-associated serine protease (MASP)-1 is an abundant component of the lectin pathway of complement. The related enzyme, MASP-2 is capable of activating the complement cascade alone. Though the concentration of MASP-1 far exceeds that of MASP-2, only a supporting role of MASP-1 has been identified regarding lectin pathway activation. Several non-complement substrates, like fibrinogen and factor XIII, have also been reported. MASP-1 belongs to the C1r/C1s/MASP family of modular serine proteases; however, its serine protease domain is evolutionary different. We have determined the crystal structure of the catalytic region of active MASP-1 and refined it to 2.55 A resolution. Unusual features of the structure are an internal salt bridge (similar to one in factor D) between the S1 Asp189 and Arg224, and a very long 60-loop. The functional and evolutionary differences between MASP-1 and the other members of the C1r/C1s/MASP family are reflected in the crystal structure. Structural comparison of the protease domains revealed that the substrate binding groove of MASP-1 is wide and resembles that of trypsin rather than early complement proteases explaining its relaxed specificity. Also, MASP-1's multifunctional behavior as both a complement and a coagulation enzyme is in accordance with our observation that antithrombin in the presence of heparin is a more potent inhibitor of MASP-1 than C1 inhibitor. Overall, MASP-1 behaves as a promiscuous protease. The structure shows that its substrate binding groove is accessible; however, its reactivity could be modulated by an unusually large 60-loop and an internal salt bridge involving the S1 Asp.
- Institute of Enzymology, Biological Research Center, Hungarian Academy of Sciences, Budapest, Hungary. dobo@enzim.hu
Organizational Affiliation: