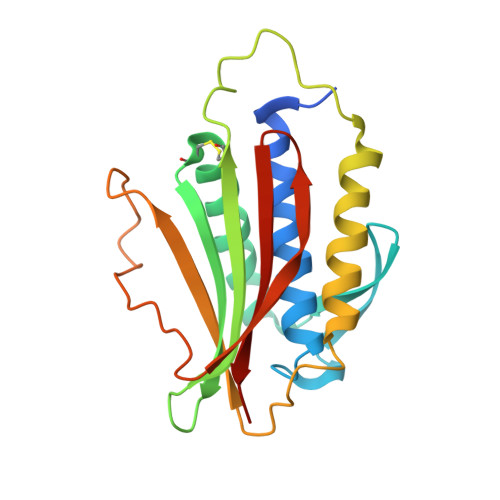
Structure of an intermediate conformer of the spindle checkpoint protein Mad2.
Hara, M., Ozkan, E., Sun, H., Yu, H., Luo, X.(2015) Proc Natl Acad Sci U S A 112: 11252-11257
- PubMed: 26305957 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1512197112
- Primary Citation Related Structures:
3GMH - PubMed Abstract:
The spindle checkpoint senses unattached kinetochores during prometaphase and inhibits the anaphase-promoting complex or cyclosome (APC/C), thus ensuring accurate chromosome segregation. The checkpoint protein mitotic arrest deficient 2 (Mad2) is an unusual protein with multiple folded states. Mad2 adopts the closed conformation (C-Mad2) in a Mad1-Mad2 core complex. In mitosis, kinetochore-bound Mad1-C-Mad2 recruits latent, open Mad2 (O-Mad2) from the cytosol and converts it to an intermediate conformer (I-Mad2), which can then bind and inhibit the APC/C activator cell division cycle 20 (Cdc20) as C-Mad2. Here, we report the crystal structure and NMR analysis of I-Mad2 bound to C-Mad2. Although I-Mad2 retains the O-Mad2 fold in crystal and in solution, its core structural elements undergo discernible rigid-body movements and more closely resemble C-Mad2. Residues exhibiting methyl chemical shift changes in I-Mad2 form a contiguous, interior network that connects its C-Mad2-binding site to the conformationally malleable C-terminal region. Mutations of residues at the I-Mad2-C-Mad2 interface hinder I-Mad2 formation and impede the structural transition of Mad2. Our study provides insight into the conformational activation of Mad2 and establishes the basis of allosteric communication between two distal sites in Mad2.
- Department of Pharmacology, University of Texas Southwestern Medical Center, Dallas, TX 75390;
Organizational Affiliation: