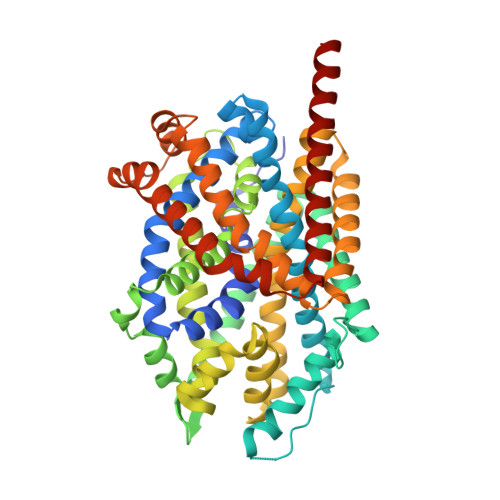
Binding of an octylglucoside detergent molecule in the second substrate (S2) site of LeuT establishes an inhibitor-bound conformation
Quick, M., Winther, A.M.L., Shi, L., Nissen, P., Weinstein, H., Javitch, J.A.(2009) Proc Natl Acad Sci U S A 106: 5563-5568
- PubMed: 19307590 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0811322106
- Primary Citation Related Structures:
3GJC, 3GJD - PubMed Abstract:
The first crystal structure of the neurotransmitter/sodium symporter homolog LeuT revealed an occluded binding pocket containing leucine and 2 Na(+); later structures showed tricyclic antidepressants (TCAs) in an extracellular vestibule approximately 11 A above the bound leucine and 2 Na(+). We recently found this region to be a second binding (S2) site and that binding of substrate to this site triggers Na(+)-coupled substrate symport. Here, we show a profound inhibitory effect of n-octyl-beta-d-glucopyranoside (OG), the detergent used for LeuT crystallization, on substrate binding to the S2 site. In parallel, we determined at 2.8 A the structure of LeuT-E290S, a mutant that, like LeuT-WT, binds 2 substrate molecules. This structure was similar to that of WT and clearly revealed an OG molecule in the S2 site. We also observed electron density at the S2 site in LeuT-WT crystals, and this also was accounted for by an OG molecule in that site. Computational analyses, based on the available crystal structures of LeuT, indicated the nature of structural arrangements in the extracellular region of LeuT that differentiate the actions of substrates from inhibitors bound in the S2 site. We conclude that the current LeuT crystal structures, all of which have been solved in OG, represent functionally blocked forms of the transporter, whereas a substrate bound in the S2 site will promote a different state that is essential for Na(+)-coupled symport.
- Center for Molecular Recognition, Columbia University College of Physicians and Surgeons, 630 West 168th Street, New York, NY 10032, USA.
Organizational Affiliation: