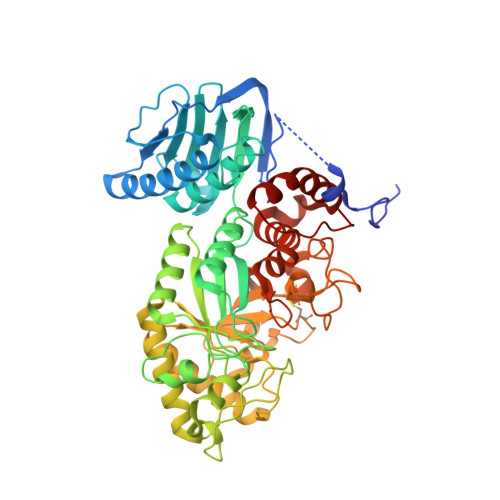
Molecular cloning and crystal structural analysis of a novel beta-N-acetylhexosaminidase from Paenibacillus sp. TS12 capable of degrading glycosphingolipids
Sumida, T., Ishii, R., Yanagisawa, T., Yokoyama, S., Ito, M.(2009) J Mol Biology 392: 87-99
- PubMed: 19524595 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.06.025
- Primary Citation Related Structures:
3GH4, 3GH5, 3GH7 - PubMed Abstract:
We report the molecular cloning and characterization of two novel beta-N-acetylhexosaminidases (beta-HEX, EC 3.2.1.52) from Paenibacillus sp. strain TS12. The two beta-HEXs (Hex1 and Hex2) were 70% identical in primary structure, and the N-terminal region of both enzymes showed significant similarity with beta-HEXs belonging to glycoside hydrolase family 20 (GH20). Interestingly, however, the C-terminal region of Hex1 and Hex2 shared no sequence similarity with the GH20 beta-HEXs or other known proteins. Both recombinant enzymes, expressed in Escherichia coli BL21(DE3), hydrolyzed the beta-N-acetylhexosamine linkage of chitooligosaccharides and glycosphingolipids such as asialo GM2 and Gb4Cer in the absence of detergent. However, the enzyme was not able to hydrolyze GM2 ganglioside in the presence or in the absence of detergent. We determined three crystal structures of Hex1; the Hex1 deletion mutant Hex1-DeltaC at a resolution of 1.8 A; Hex1-DeltaC in complex with beta-N-acetylglucosamine at 1.6 A; and Hex1-DeltaC in complex with beta-N-acetylgalactosamine at 1.9 A. We made a docking model of Hex1-DeltaC with GM2 oligosaccharide, revealing that the sialic acid residue of GM2 could hinder access of the substrate to the active site cavity. This is the first report describing the molecular cloning, characterization and X-ray structure of a procaryotic beta-HEX capable of hydrolyzing glycosphingolipids.
- Department of Bioscience and Biotechnology, Kyushu University, Hakozaki, Higashi-ku, Fukuoka, Japan.
Organizational Affiliation: