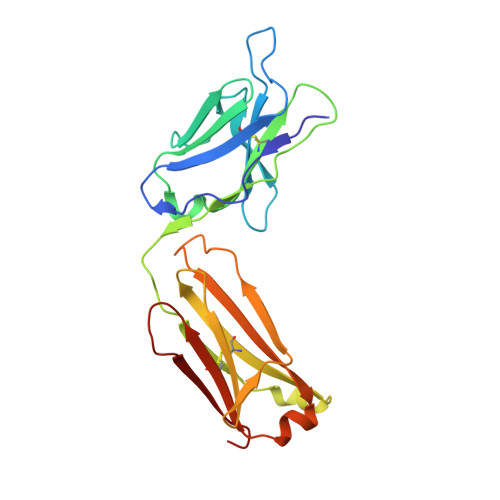
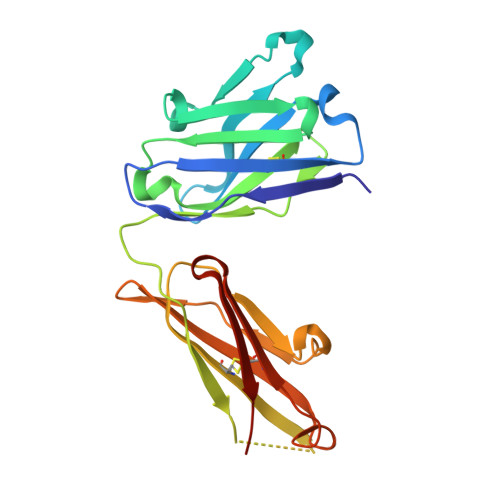
Structural Mimicry of O-Antigen by a Peptide Revealed in a Complex with an Antibody Raised against Shigella flexneri Serotype 2a
Theillet, F.-X., Saul, F.A., Vulliez-Le Normand, B., Hoos, S., Felici, F., Weintraub, A., Mulard, L.A., Phalipon, A., Delepierre, M., Bentley, G.A.(2009) J Mol Biology 388: 839-850
- PubMed: 19328810 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.03.057
- Primary Citation Related Structures:
3GGW - PubMed Abstract:
The use of carbohydrate-mimicking peptides to induce immune responses against surface polysaccharides of pathogenic bacteria offers a novel approach to vaccine development. Factors governing antigenic and immunogenic mimicry, however, are complex and poorly understood. We have addressed this question using the anti-lipopolysaccharide monoclonal antibody F22-4, which was raised against Shigella flexneri serotype 2a and shown to protect against homologous infection in a mouse model. In a previous crystallographic study, we described F22-4 in complex with two synthetic fragments of the O-antigen, the serotype-specific saccharide moiety of lipopolysaccharide. Here, we present a crystallographic and NMR study of the interaction of F22-4 with a dodecapeptide selected by phage display using the monoclonal antibody. Like the synthetic decasaccharide, the peptide binds to F22-4 with micromolar affinity. Although the peptide and decasaccharide use very similar regions of the antigen-binding site, indicating good antigenic mimicry, immunogenic mimicry by the peptide was not observed. The F22-4-antigen interaction is significantly more hydrophobic with the peptide than with oligosaccharides; nonetheless, all hydrogen bonds formed between the peptide and F22-4 have equivalents in the oligosaccharide complex. Two bridging water molecules are also in common, adding to partial structural mimicry. Whereas the bound peptide is entirely helical, its structure in solution, as shown by NMR, is helical in the central region only. Moreover, docking the NMR structure into the antigen-binding site shows that steric hindrance would occur, revealing poor complementarity between the major solution conformation and the antibody that could contribute to the absence of immunogenic mimicry.
- Institut Pasteur, Unité de RMN des Biomolécules, CNRS URA 2185, Paris, France.
Organizational Affiliation: