Characterization of a Bacillus subtilis transporter for petrobactin, an anthrax stealth siderophore.
Zawadzka, A.M., Kim, Y., Maltseva, N., Nichiporuk, R., Fan, Y., Joachimiak, A., Raymond, K.N.(2009) Proc Natl Acad Sci U S A 106: 21854-21859
- PubMed: 19955416 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0904793106
- Primary Citation Related Structures:
3GFV - PubMed Abstract:
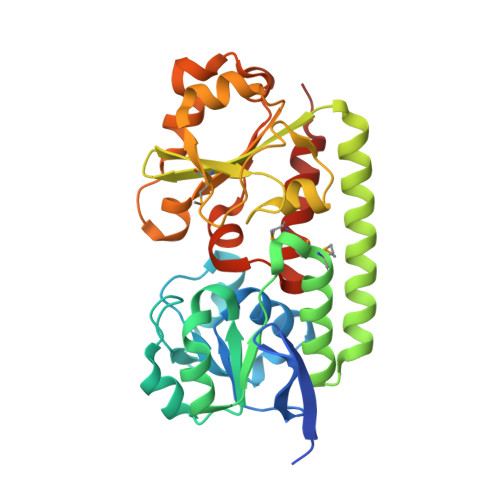
Iron deprivation activates the expression of components of the siderophore-mediated iron acquisition systems in Bacillus subtilis, including not only the synthesis and uptake of its siderophore bacillibactin but also expression of multiple ABC transporters for iron scavenging using xenosiderophores. The yclNOPQ operon is shown to encode the complete transporter for petrobactin (PB), a photoreactive 3,4-catecholate siderophore produced by many members of the B. cereus group, including B. anthracis. Isogenic disruption mutants in the yclNOPQ transporter, including permease YclN, ATPase YclP, and a substrate-binding protein YclQ, are unable to use either PB or the photoproduct of FePB (FePB(nu)) for iron delivery and growth, in contrast to the wild-type B. subtilis. Complementation of the mutations with the copies of the respective genes restores this capability. The YclQ receptor binds selectively iron-free and ferric PB, the PB precursor, 3,4-dihydroxybenzoic acid (3,4-DHB), and FePB(nu) with high affinity; the ferric complexes are seen in ESI-MS, implying strong electrostatic interaction between the protein-binding pocket and siderophore. The first structure of a gram-positive siderophore receptor is presented. The 1.75-A crystal structure of YclQ reveals a bilobal periplasmic binding protein (PBP) fold consisting of two alpha/beta/alpha sandwich domains connected by a long alpha-helix with the binding pocket containing conserved positively charged and aromatic residues and large enough to accommodate FePB. Orthologs of the B. subtilis PB-transporter YclNOPQ in PB-producing Bacilli are likely contributors to the pathogenicity of these species and provide a potential target for antibacterial strategies.
- Department of Chemistry, University of California, Berkeley, CA 94720-1460, USA.
Organizational Affiliation: