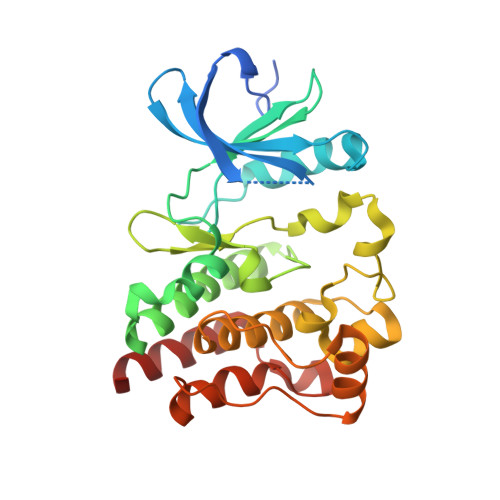
Structures of human Bruton's tyrosine kinase in active and inactive conformations suggest a mechanism of activation for TEC family kinases.
Marcotte, D.J., Liu, Y.T., Arduini, R.M., Hession, C.A., Miatkowski, K., Wildes, C.P., Cullen, P.F., Hong, V., Hopkins, B.T., Mertsching, E., Jenkins, T.J., Romanowski, M.J., Baker, D.P., Silvian, L.F.(2010) Protein Sci 19: 429-439
- PubMed: 20052711 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.321
- Primary Citation Related Structures:
3GEN, 3K54 - PubMed Abstract:
Bruton's tyrosine kinase (BTK), a member of the TEC family of kinases, plays a crucial role in B-cell maturation and mast cell activation. Although the structures of the unphosphorylated mouse BTK kinase domain and the unphosphorylated and phosphorylated kinase domains of human ITK are known, understanding the kinase selectivity profiles of BTK inhibitors has been hampered by the lack of availability of a high resolution, ligand-bound BTK structure. Here, we report the crystal structures of the human BTK kinase domain bound to either Dasatinib (BMS-354825) at 1.9 A resolution or to 4-amino-5-(4-phenoxyphenyl)-7H-pyrrolospyrimidin- 7-yl-cyclopentane at 1.6 A resolution. This data provides information relevant to the development of small molecule inhibitors targeting BTK and the TEC family of nonreceptor tyrosine kinases. Analysis of the structural differences between the TEC and Src families of kinases near the Trp-Glu-Ile motif in the N-terminal region of the kinase domain suggests a mechanism of regulation of the TEC family members.
- Biogen Idec, Inc., Drug Discovery Department, 12 Cambridge Center, Cambridge, Massachusetts 02142, USA.
Organizational Affiliation: