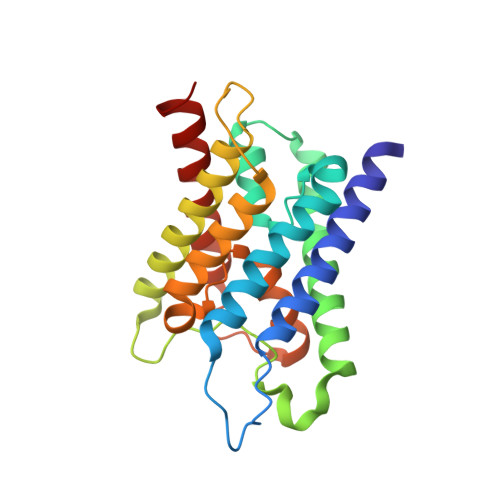
Crystal structure of human aquaporin 4 at 1.8 A and its mechanism of conductance.
Ho, J.D., Yeh, R., Sandstrom, A., Chorny, I., Harries, W.E., Robbins, R.A., Miercke, L.J., Stroud, R.M.(2009) Proc Natl Acad Sci U S A 106: 7437-7442
- PubMed: 19383790 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0902725106
- Primary Citation Related Structures:
3GD8 - PubMed Abstract:
Aquaporin (AQP) 4 is the predominant water channel in the mammalian brain, abundantly expressed in the blood-brain and brain-cerebrospinal fluid interfaces of glial cells. Its function in cerebral water balance has implications in neuropathological disorders, including brain edema, stroke, and head injuries. The 1.8-A crystal structure reveals the molecular basis for the water selectivity of the channel. Unlike the case in the structures of water-selective AQPs AqpZ and AQP1, the asparagines of the 2 Asn-Pro-Ala motifs do not hydrogen bond to the same water molecule; instead, they bond to 2 different water molecules in the center of the channel. Molecular dynamics simulations were performed to ask how this observation bears on the proposed mechanisms for how AQPs remain totally insulating to any proton conductance while maintaining a single file of hydrogen bonded water molecules throughout the channel.
- Graduate Program in Chemistry and Chemical Biology and Department of Biochemistry and Biophysics, Genentech Hall, University of California, 600 16th Street, San Francisco, CA 94158-2517, USA.
Organizational Affiliation: