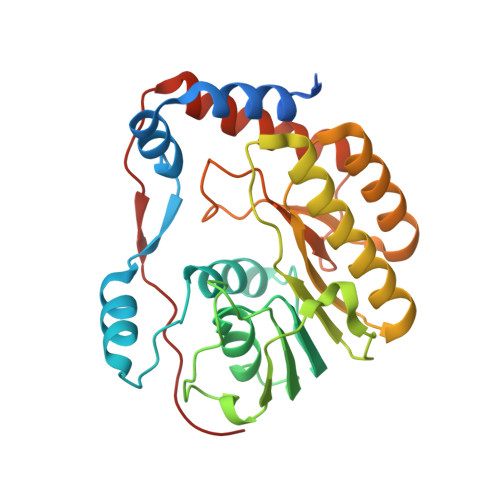
Crystal structure of a methyltransferase from a no-known-vector Flavivirus
Bollati, M., Milani, M., Mastrangelo, E., de Lambellarie, X., Canard, B., Bolognesi, M.(2009) Biochem Biophys Res Commun 382: 200-204
- PubMed: 19275894 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2009.03.008
- Primary Citation Related Structures:
3GCZ - PubMed Abstract:
Presently known flaviviruses belong to three major evolutionary branches: tick-borne viruses, mosquito-borne viruses and viruses with no known vector. Here we present the crystal structure of the Yokose virus methyltransferase at 1.7A resolution, the first structure of a methyltransferase of a Flavivirus with no known vector. Structural comparison of three methyltransferases representative of each of the Flavivirus branches shows that fold and structures are closely conserved, most differences being related to surface loops flexibility. Analysis of the conserved residues throughout all the sequenced flaviviral methyltransferases reveals that, besides the central cleft hosting the substrate and cofactor binding sites, a second, almost continuous, patch is conserved and points away from active site towards the back of the protein. The high level of structural conservation in this region could be functional for the methyltransferase/RNA interaction and stabilization of the ensuing complex.
- Department of Biomolecular Sciences and Biotechnology, CNR-INFM and CIMAINA, University of Milano, Via Celoria 26, 20133 Milano, Italy.
Organizational Affiliation: