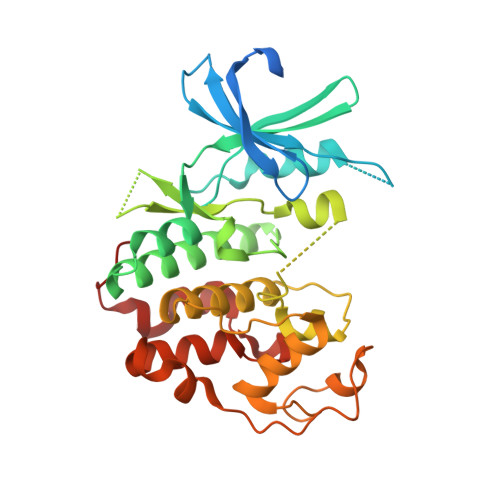
Structure of a cyclin-dependent kinase from Giardia lamblia.
Leibly, D.J., Newling, P.A., Abendroth, J., Guo, W., Kelley, A., Stewart, L.J., Van Voorhis, W.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 1084-1089
- PubMed: 21904054 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111018070
- Primary Citation Related Structures:
3GBZ, 3GC0 - PubMed Abstract:
Giardia lamblia is the etiologic agent of giardiasis, a water-borne infection that is prevalent throughout the world. The need for new therapeutics for the treatment of giardiasis is of paramount importance. Owing to the ubiquitous nature of kinases and their vital importance in organisms, they are potential drug targets. In this paper, the first structure of a cyclin-dependent kinase (CDK) from G. lamblia (GlCDK; UniProt A8BZ95) is presented. CDKs are cell-cycle-associated kinases that are actively being pursued as targets for anticancer drugs as well as for antiparasitic chemotherapy. Generally, a CDK forms a complex with its associated cyclin. This CDK-cyclin complex is active and acts as a serine/threonine protein kinase. Typically, CDKs are responsible for the transition to the next phase of the cell cycle. Although the structure of GlCDK with its associated cyclin was not solved, the 1.85 Å resolution structure of apo GlCDK and a 2.0 Å resolution structure of GlCDK in complex with adenosine monophosphate are presented and the structural differences from the orthologous human CDK2 and CDK3 are discussed.
- Seattle Structural Genomics Center for Infectious Disease (SSGCID), USA.
Organizational Affiliation: