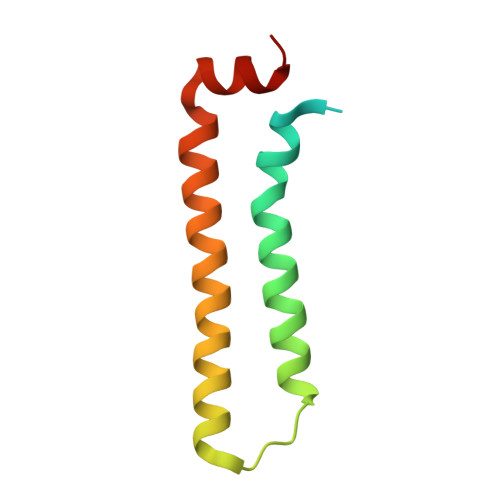
Structure of the RNA-Binding Domain of Nodamura Virus Protein B2, a Suppressor of RNA Interference.
Korber, S., Shaik Syed Ali, P., Chen, J.C.(2009) Biochemistry 48: 2307-2309
- PubMed: 19249868 Search on PubMed
- DOI: https://doi.org/10.1021/bi900126s
- Primary Citation Related Structures:
3G80 - PubMed Abstract:
Protein B2 from Nodamura virus (NMV B2), a member of the Nodavirus family, acts as a suppressor of RNA interference (RNAi). The N-terminal domain of NMV B2, consisting of residues 1-79, recognizes double-stranded RNA (dsRNA). The 2.5 A crystal structure of the RNA-binding domain of NMV B2 shows a dimeric, helical bundle structure. The structure shows a conserved set of RNA-binding residues compared with flock house virus B2, despite limited sequence identity. The crystal packing places the RNA-binding residues along one face of symmetry-related molecules, suggesting a potential platform for recognition of dsRNA.
- Institute of Biophysical Chemistry, Goethe University Frankfurt, Max-von-Laue-Strasse 9, 60438 Frankfurt, Germany.
Organizational Affiliation: