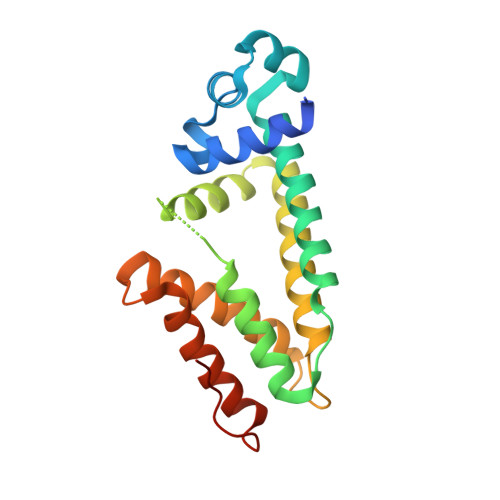
Structure and function of the macrolide biosensor protein, MphR(A), with and without erythromycin
Zheng, J., Sagar, V., Smolinsky, A., Bourke, C., LaRonde-LeBlanc, N., Cropp, T.A.(2009) J Mol Biology 387: 1250-1260
- PubMed: 19265703 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.02.058
- Primary Citation Related Structures:
3FRQ, 3G56 - PubMed Abstract:
The regulatory protein MphR(A) has recently seen extensive use in synthetic biological applications, such as metabolite sensing and exogenous control of gene expression. This protein negatively regulates the expression of a macrolide 2'-phosphotransferase I resistance gene (mphA) via binding to a 35-bp DNA operator upstream of the start codon and is de-repressed by the presence of erythromycin. Here, we present the refined crystal structure of the MphR(A) protein free of erythromycin and that of the MphR(A) protein with bound erythromycin at 2.00- and 1.76-A resolutions, respectively. We also studied the DNA binding properties of the protein and identified mutants of MphR(A) that are defective in gene repression and ligand binding in a cell-based reporter assay. The combination of these two structures illustrates the molecular basis of erythromycin-induced gene expression and provides a framework for additional applied uses of this protein in the isolation and engineered biosynthesis of polyketide natural products.
- Department of Chemistry and Biochemistry, University of Maryland, College Park, MD 20742, USA.
Organizational Affiliation: