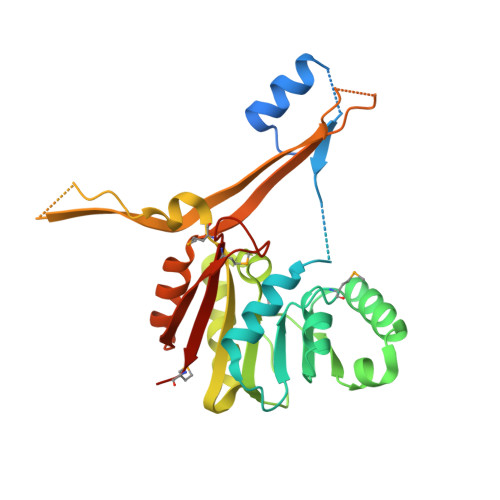
Structure and function of the glycopeptide N-methyltransferase MtfA, a tool for the biosynthesis of modified glycopeptide antibiotics.
Shi, R., Lamb, S.S., Zakeri, B., Proteau, A., Cui, Q., Sulea, T., Matte, A., Wright, G.D., Cygler, M.(2009) Chem Biol 16: 401-410
- PubMed: 19389626 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2009.02.007
- Primary Citation Related Structures:
3G2M, 3G2O, 3G2P, 3G2Q - PubMed Abstract:
There is a considerable interest in the modification of existing antibiotics to generate new antimicrobials. Glycopeptide antibiotics (GPAs) are effective against serious Gram-positive bacterial pathogens including methicillin-resistant Staphylococcus aureus. However, resistance to these antibiotics is becoming a serious problem requiring new strategies. We show that the Amycolatopsis orientalis (S)-adenosyl-L-methionine-dependent methyltransferase MtfA, from the vancomycin-class GPA chloroeremomycin biosynthetic pathway, catalyzes in vivo and in vitro methyl transfer to generate methylated GPA derivatives of the teicoplanin class. The crystal structure of MtfA complexed with (S)-adenosyl-L-methionine, (S)-adenosylhomocysteine, or sinefungin inhibitor, coupled with mutagenesis, identified His228 as a likely general base required for methyl transfer to the N terminus of the glycopeptide. Computational docking and molecular dynamics simulations were used to model binding of demethyl-vancomycin aglycone to MtfA. These results demonstrate its utility as a tool for engineering methylated analogs of GPAs.
- Department of Biochemistry, McGill University, 3655 Promenade Sir William Osler, Montréal, QC H3G 1Y6, Canada.
Organizational Affiliation: