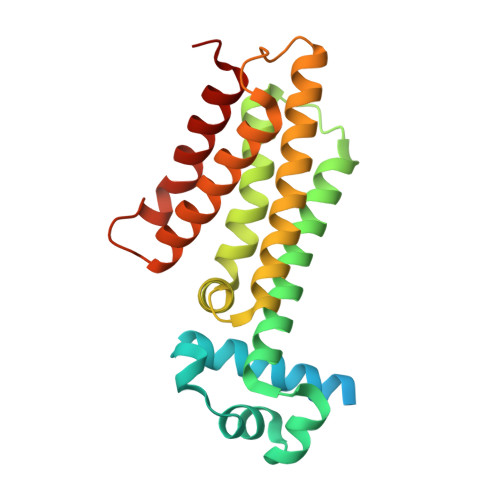
Synthetic EthR inhibitors boost antituberculous activity of ethionamide.
Willand, N., Dirie, B., Carette, X., Bifani, P., Singhal, A., Desroses, M., Leroux, F., Willery, E., Mathys, V., Deprez-Poulain, R., Delcroix, G., Frenois, F., Aumercier, M., Locht, C., Villeret, V., Deprez, B., Baulard, A.R.(2009) Nat Med 15: 537-544
- PubMed: 19412174 Search on PubMed
- DOI: https://doi.org/10.1038/nm.1950
- Primary Citation Related Structures:
3G1L, 3G1M, 3G1O - PubMed Abstract:
The side effects associated with tuberculosis therapy bring with them the risk of noncompliance and subsequent drug resistance. Increasing the therapeutic index of antituberculosis drugs should thus improve treatment effectiveness. Several antituberculosis compounds require in situ metabolic activation to become inhibitory. Various thiocarbamide-containing drugs, including ethionamide, are activated by the mycobacterial monooxygenase EthA, the production of which is controlled by the transcriptional repressor EthR. Here we identify drug-like inhibitors of EthR that boost the bioactivation of ethionamide. Compounds designed and screened for their capacity to inhibit EthR-DNA interaction were co-crystallized with EthR. We exploited the three-dimensional structures of the complexes for the synthesis of improved analogs that boosted the ethionamide potency in culture more than tenfold. In Mycobacterium tuberculosis-infected mice, one of these analogs, BDM31343, enabled a substantially reduced dose of ethionamide to lessen the mycobacterial load as efficiently as the conventional higher-dose treatment. This provides proof of concept that inhibiting EthR improves the therapeutic index of thiocarbamide derivatives, which should prompt reconsideration of their use as first-line drugs.
- Institut National de la Santé et de la Recherche Médicale, Lille, France.
Organizational Affiliation: