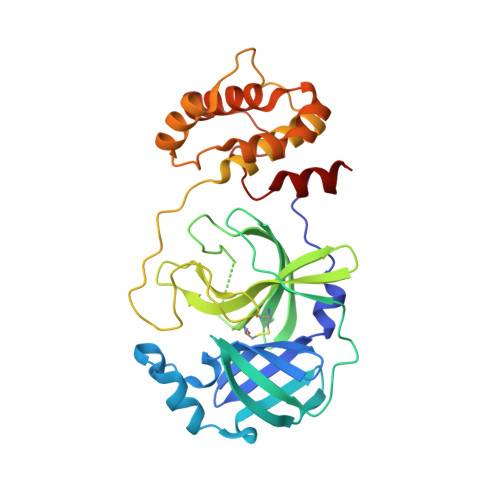
Mutation of Asn28 disrupts the dimerization and enzymatic activity of SARS 3CL(pro) .
Barrila, J., Gabelli, S.B., Bacha, U., Amzel, L.M., Freire, E.(2010) Biochemistry 49: 4308-4317
- PubMed: 20420403 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi1002585
- Primary Citation Related Structures:
3FZD - PubMed Abstract:
Coronaviruses are responsible for a significant proportion of annual respiratory and enteric infections in humans and other mammals. The most prominent of these viruses is the severe acute respiratory syndrome coronavirus (SARS-CoV) which causes acute respiratory and gastrointestinal infection in humans. The coronavirus main protease, 3CL(pro), is a key target for broad-spectrum antiviral development because of its critical role in viral maturation and high degree of structural conservation among coronaviruses. Dimerization is an indispensable requirement for the function of SARS 3CL(pro) and is regulated through mechanisms involving both direct and long-range interactions in the enzyme. While many of the binding interactions at the dimerization interface have been extensively studied, those that are important for long-range control are not well-understood. Characterization of these dimerization mechanisms is important for the structure-based design of new treatments targeting coronavirus-based infections. Here we report that Asn28, a residue 11 A from the closest residue in the opposing monomer, is essential for the enzymatic activity and dimerization of SARS 3CL(pro). Mutation of this residue to alanine almost completely inactivates the enzyme and results in a 19.2-fold decrease in the dimerization K(d). The crystallographic structure of the N28A mutant determined at 2.35 A resolution reveals the critical role of Asn28 in maintaining the structural integrity of the active site and in orienting key residues involved in binding at the dimer interface and substrate catalysis. These findings provide deeper insight into complex mechanisms regulating the activity and dimerization of SARS 3CL(pro).
- Department of Biology, Johns Hopkins University, Baltimore, Maryland 21218, USA.
Organizational Affiliation: