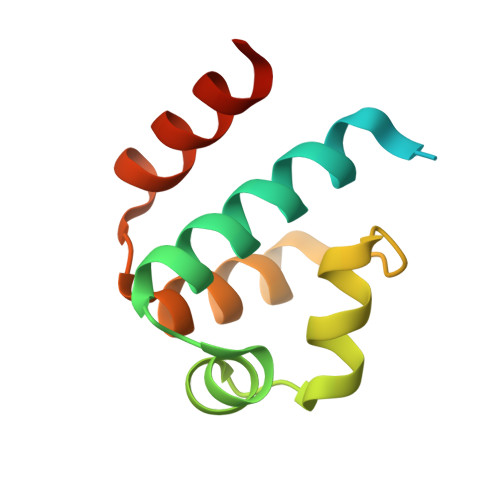
Structure of the restriction-modification controller protein C.Esp1396I.
Ball, N., Streeter, S.D., Kneale, G.G., McGeehan, J.E.(2009) Acta Crystallogr D Biol Crystallogr 65: 900-905
- PubMed: 19690367 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444909020514
- Primary Citation Related Structures:
3FYA, 3G5G - PubMed Abstract:
The controller protein of the Esp1396I restriction-modification (R-M) system binds differentially to three distinct operator sequences upstream of the methyltransferase (M) and endonuclease (R) genes to regulate the timing of gene expression. The crystal structure of a complex of the protein with two adjacent operator DNA sequences has been reported; however, the structure of the free protein has not yet been determined. Here, the crystal structure of the free protein is reported, with seven dimers in the asymmetric unit. Two of the 14 monomers show an alternative conformation to the major conformer in which the side chains of residues 43-46 in the loop region flanking the DNA-recognition helix are displaced by up to 10 A. It is proposed that the adoption of these two conformational states may play a role in DNA-sequence promiscuity. The two alternative conformations are also found in the R35A mutant structure, which is otherwise identical to the native protein. Comparison of the free and bound protein structures shows a 1.4 A displacement of the recognition helices when the dimer is bound to its DNA target.
- Biophysics Laboratories, Institute of Biomedical and Biomolecular Sciences, School of Biological Sciences, University of Portsmouth, Portsmouth, UK.
Organizational Affiliation: