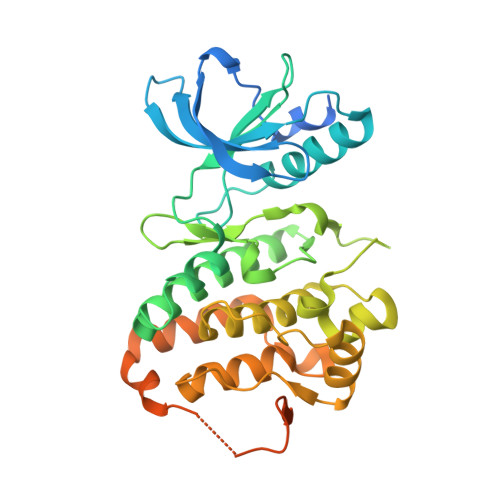
Structural recognition of an optimized substrate for the ephrin family of receptor tyrosine kinases.
Davis, T.L., Walker, J.R., Allali-Hassani, A., Parker, S.A., Turk, B.E., Dhe-Paganon, S.(2009) FEBS J 276: 4395-4404
- PubMed: 19678838 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/j.1742-4658.2009.07147.x
- Primary Citation Related Structures:
3FXX, 3FY2 - PubMed Abstract:
Ephrin receptor tyrosine kinase A3 (EphA3, EC 2.7.10.1) is a member of a unique branch of the kinome in which downstream signaling occurs in both ligand- and receptor-expressing cells. Consequently, the ephrins and ephrin receptor tyrosine kinases often mediate processes involving cell-cell contact, including cellular adhesion or repulsion, developmental remodeling and neuronal mapping. The receptor is also frequently overexpressed in invasive cancers, including breast, small-cell lung and gastrointestinal cancers. However, little is known about direct substrates of EphA3 kinase and no chemical probes are available. Using a library approach, we found a short peptide sequence that is a good substrate for EphA3 and is suitable for co-crystallization studies. Complex structures show multiple contacts between kinase and substrates; in particular, two residues undergo conformational changes and by mutation are found to be important for substrate binding and turnover. In addition, a difference in catalytic efficiency between EPH kinase family members is observed. These results provide insight into the mechanism of substrate binding to these developmentally integral enzymes.
- Structural Genomics Consortium, University of Toronto, Canada.
Organizational Affiliation: