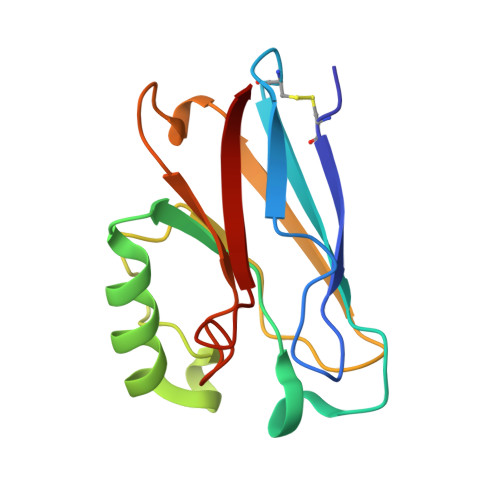
Metal-binding loop length and not sequence dictates structure.
Sato, K., Li, C., Salard, I., Thompson, A.J., Banfield, M.J., Dennison, C.(2009) Proc Natl Acad Sci U S A 106: 5616-5621
- PubMed: 19299503 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0811324106
- Primary Citation Related Structures:
3FS9, 3FSA, 3FSV, 3FSW, 3FSZ, 3FT0 - PubMed Abstract:
The C-terminal copper-binding loop in the beta-barrel fold of the cupredoxin azurin has been replaced with a range of sequences containing alanine, glycine, and valine residues to assess the importance of amino acid composition and the length of this region. The introduction of 2 and 4 alanines between the coordinating Cys, His, and Met results in loop structures matching those in naturally occurring proteins with the same loop lengths. A loop with 4 alanines between the Cys and His and 3 between the His and Met ligands has a structure identical to that of the WT protein, whose loop is the same length. Loop structure is dictated by length and not sequence allowing the properties of the main surface patch for interactions with partners, to which the loop is a major contributor, to be optimized. Loops with 2 amino acids between the ligands using glycine, alanine, and valine residues have been compared. An empirical relationship is found between copper site protection by the loop and reduction potential. A loop adorned with 4 methyl groups is sufficient to protect the copper ion, enabling most sequences to adequately perform this task. The mutant with 3 alanine residues between the ligands forms a strand-swapped dimer in the crystal structure, an arrangement that has not, to our knowledge, been seen previously for this family of proteins. Cupredoxins function as redox shuttles and are required to be monomeric; therefore, none have evolved with a metal-binding loop of this length.
- Institute for Cell and Molecular Biosciences, Medical School, Newcastle University, Newcastle upon Tyne NE2 4HH, United Kingdom.
Organizational Affiliation: