Phosphorylated self-peptides alter human leukocyte antigen class I-restricted antigen presentation and generate tumor-specific epitopes
Petersen, J., Wurzbacher, S.J., Williamson, N.A., Ramarathinam, S.H., Reid, H.H., Nair, A.K., Zhao, A.Y., Nastovska, R., Rudge, G., Rossjohn, J., Purcell, A.W.(2009) Proc Natl Acad Sci U S A 106: 2776-2781
- PubMed: 19196958 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0812901106
- Primary Citation Related Structures:
3FQN, 3FQR, 3FQT, 3FQU, 3FQW, 3FQX - PubMed Abstract:
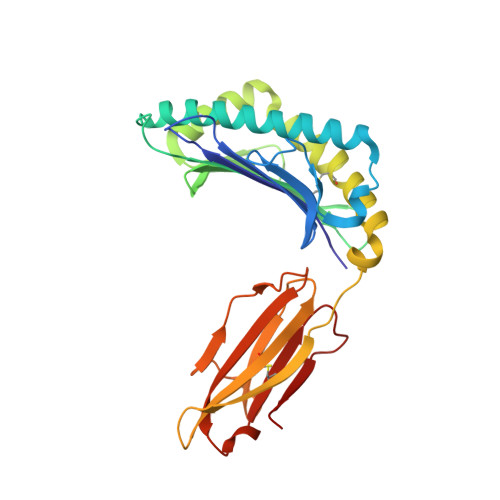
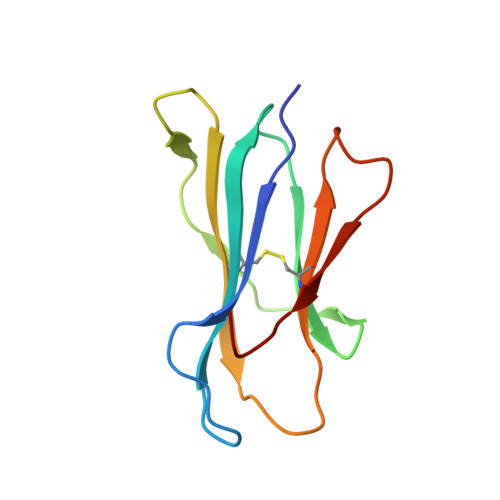
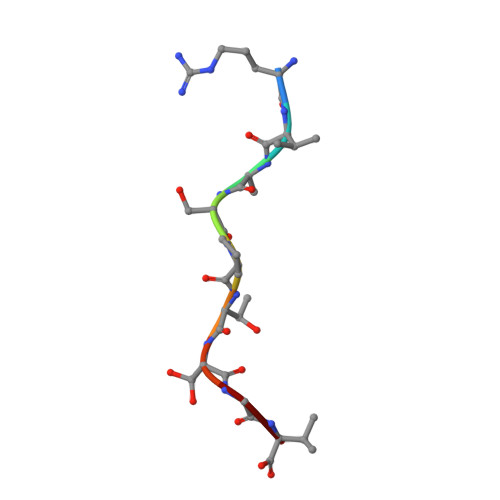
Human leukocyte antigen (HLA) class I molecules present a variety of posttranslationally modified epitopes at the cell surface, although the consequences of such presentation remain largely unclear. Phosphorylation plays a critical cellular role, and deregulation in phosphate metabolism is associated with disease, including autoimmunity and tumor immunity. We have solved the high-resolution structures of 3 HLA A2-restricted phosphopeptides associated with tumor immunity and compared them with the structures of their nonphosphorylated counterparts. Phosphorylation of the epitope was observed to affect the structure and mobility of the bound epitope. In addition, the phosphoamino acid stabilized the HLA peptide complex in an epitope-specific manner and was observed to exhibit discrete flexibility within the antigen-binding cleft. Collectively, our data suggest that phosphorylation generates neoepitopes that represent demanding targets for T-cell receptor ligation. These findings provide insights into the mode of phosphopeptide presentation by HLA as well as providing a platform for the rational design of a generation of posttranslationally modified tumor vaccines.
- Protein Crystallography Unit, Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Victoria 3800, Australia.
Organizational Affiliation: