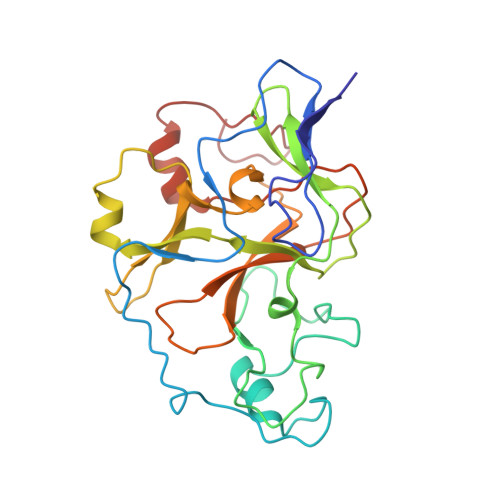
Structural basis for G9a-like protein lysine methyltransferase inhibition by BIX-01294.
Chang, Y., Zhang, X., Horton, J.R., Upadhyay, A.K., Spannhoff, A., Liu, J., Snyder, J.P., Bedford, M.T., Cheng, X.(2009) Nat Struct Mol Biol 16: 312-317
- PubMed: 19219047 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1560
- Primary Citation Related Structures:
3FPD - PubMed Abstract:
Histone lysine methylation is an important epigenetic mark that regulates gene expression and chromatin organization. G9a and G9a-like protein (GLP) are euchromatin-associated methyltransferases that repress transcription by methylating histone H3 Lys9. BIX-01294 was originally identified as a G9a inhibitor during a chemical library screen of small molecules and has previously been used in the generation of induced pluripotent stem cells. Here we present the crystal structure of the catalytic SET domain of GLP in complex with BIX-01294 and S-adenosyl-L-homocysteine. The inhibitor is bound in the substrate peptide groove at the location where the histone H3 residues N-terminal to the target lysine lie in the previously solved structure of the complex with histone peptide. The inhibitor resembles the bound conformation of histone H3 Lys4 to Arg8, and is positioned in place by residues specific for G9a and GLP through specific interactions.
- Department of Biochemistry, Emory University School of Medicine, 1510 Clifton Road, Atlanta, Georgia 30322, USA.
Organizational Affiliation: