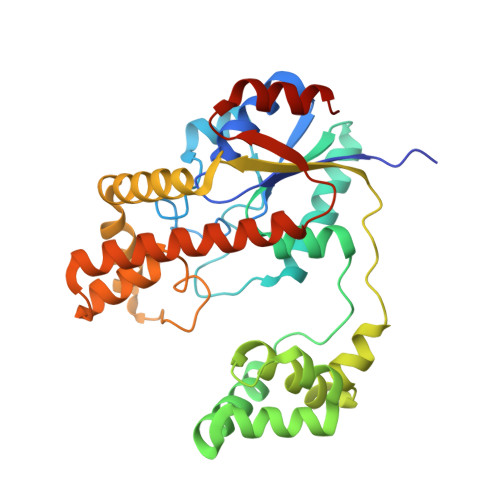
RNA-Protein Mutually Induced Fit: STRUCTURE OF ESCHERICHIA COLI ISOPENTENYL-tRNA TRANSFERASE IN COMPLEX WITH tRNA(Phe).
Seif, E., Hallberg, B.M.(2009) J Biological Chem 284: 6600-6604
- PubMed: 19158097 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.C800235200
- Primary Citation Related Structures:
3FOZ - PubMed Abstract:
tRNAs that read codons starting with U are usually modified at their A37 by isopentenyl-tRNA transferases to minimize peptidyl-tRNA slippage in translation. The consensus substrate requirements of the isopentenyl-tRNA transferase of Escherichia coli, MiaA, have been the focus of extensive study. However, the molecular basis of tRNA-MiaA recognition remains unknown. Here we describe the 2.5A crystal structure of MiaA in complex with substrate tRNA(Phe). Comparative structural analysis reveals that the enzymatic reaction involves an RNA-protein mutually induced fit mechanism in which large domain movements in MiaA provoke the partial unfolding of the substrate tRNA anticodon loop. In addition, we show how substrate tRNAs are recognized by MiaA through a combination of direct and indirect sequence readouts.
- Department of Cell and Molecular Biology, Medical Nobel Institute, Karolinska Institutet, SE-171 77 Stockholm, Sweden.
Organizational Affiliation: