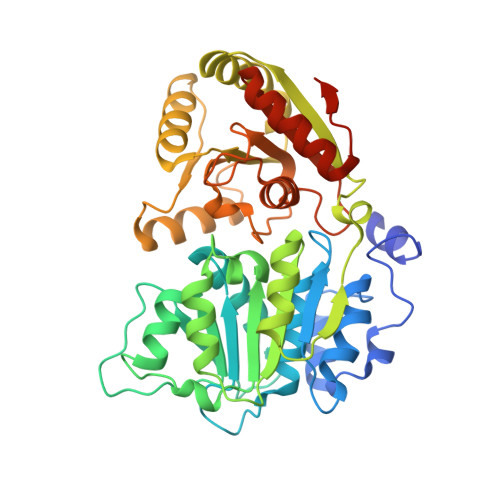
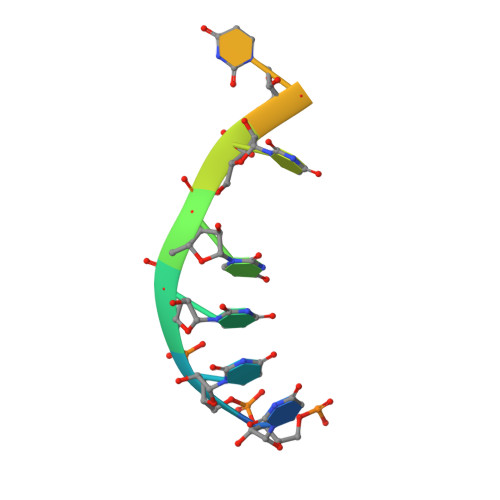
The mRNA export protein DBP5 binds RNA and the cytoplasmic nucleoporin NUP214 in a mutually exclusive manner
von Moeller, H., Basquin, C., Conti, E.(2009) Nat Struct Mol Biol 16: 247-254
- PubMed: 19219046 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.1561
- Primary Citation Related Structures:
3FHC, 3FHT - PubMed Abstract:
The DEAD-box protein DBP5 is essential for mRNA export in both yeast and humans. It binds RNA and is concentrated and locally activated at the cytoplasmic side of the nuclear pore complex. We have determined the crystal structures of human DBP5 bound to RNA and AMPPNP, and bound to the cytoplasmic nucleoporin NUP214. The structures reveal that binding of DBP5 to nucleic acid and to NUP214 is mutually exclusive. Using in vitro assays, we demonstrate that NUP214 decreases both the RNA binding and ATPase activities of DBP5. The interactions are mediated by conserved residues, implying a conserved recognition mechanism. These results suggest a framework for the consecutive steps leading to the release of mRNA at the final stages of nuclear export. More generally, they provide a paradigm for how binding of regulators can specifically inhibit DEAD-box proteins.
- Max-Planck-Institute of Biochemistry, Department of Structural Cell Biology, Am Klopferspitz 18, D-82152 Martinsried, Germany.
Organizational Affiliation: