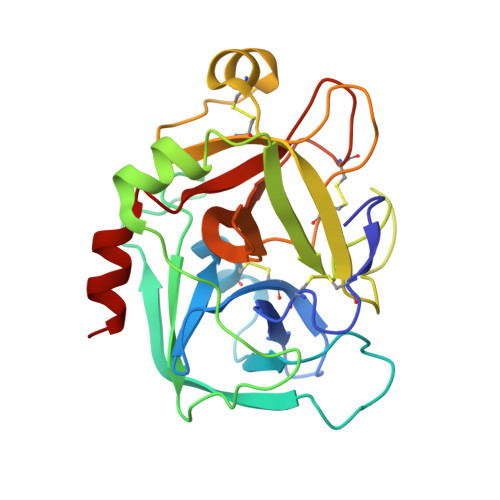
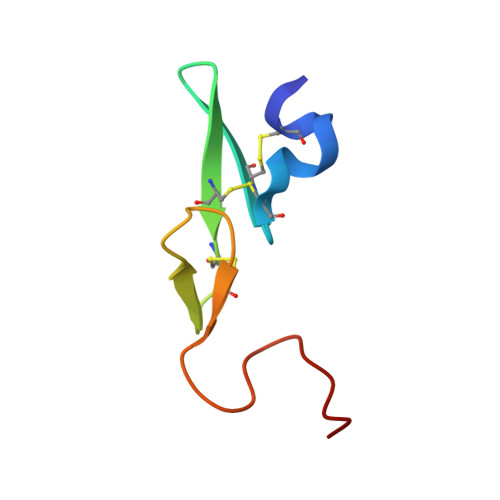
Phenyltriazolinones as potent factor Xa inhibitors.
Quan, M.L., Pinto, D.J., Rossi, K.A., Sheriff, S., Alexander, R.S., Amparo, E., Kish, K., Knabb, R.M., Luettgen, J.M., Morin, P., Smallwood, A., Woerner, F.J., Wexler, R.R.(2010) Bioorg Med Chem Lett 20: 1373-1377
- PubMed: 20100660 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2010.01.011
- Primary Citation Related Structures:
3FFG, 3KQB, 3KQC, 3KQD, 3KQE - PubMed Abstract:
We have discovered that phenyltriazolinone is a novel and potent P1 moiety for coagulation factor Xa. X-ray structures of the inhibitors with a phenyltriazolinone in the P1 position revealed that the side chain of Asp189 has reoriented resulting in a novel S1 binding pocket which is larger in size to accommodate the phenyltriazolinone P1 substrate.
- Research and Development, Bristol-Myers Squibb, PO Box 5400, Princeton, NJ 08543-5400, USA. mimi.quan@bms.com
Organizational Affiliation: