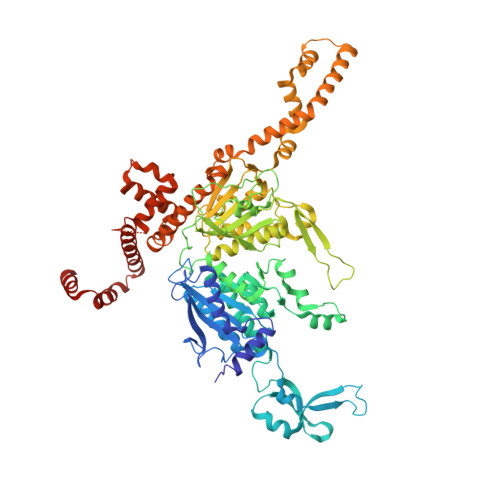
The human ACC2 CT-domain C-terminus is required for full functionality and has a novel twist.
Madauss, K.P., Burkhart, W.A., Consler, T.G., Cowan, D.J., Gottschalk, W.K., Miller, A.B., Short, S.A., Tran, T.B., Williams, S.P.(2009) Acta Crystallogr D Biol Crystallogr 65: 449-461
- PubMed: 19390150 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444909008014
- Primary Citation Related Structures:
3FF6 - PubMed Abstract:
Inhibition of acetyl-CoA carboxylase (ACC) may prevent lipid-induced insulin resistance and type 2 diabetes, making the enzyme an attractive pharmaceutical target. Although the enzyme is highly conserved amongst animals, only the yeast enzyme structure is available for rational drug design. The use of biophysical assays has permitted the identification of a specific C-terminal truncation of the 826-residue human ACC2 carboxyl transferase (CT) domain that is both functionally competent to bind inhibitors and crystallizes in their presence. This C-terminal truncation led to the determination of the human ACC2 CT domain-CP-640186 complex crystal structure, which revealed distinctions from the yeast-enzyme complex. The human ACC2 CT-domain C-terminus is comprised of three intertwined alpha-helices that extend outwards from the enzyme on the opposite side to the ligand-binding site. Differences in the observed inhibitor conformation between the yeast and human structures are caused by differing residues in the binding pocket.
- Department of Computational and Structural Chemistry, GlaxoSmithKline Inc., Five Moore Drive, Research Triangle Park, NC 27709, USA.
Organizational Affiliation: