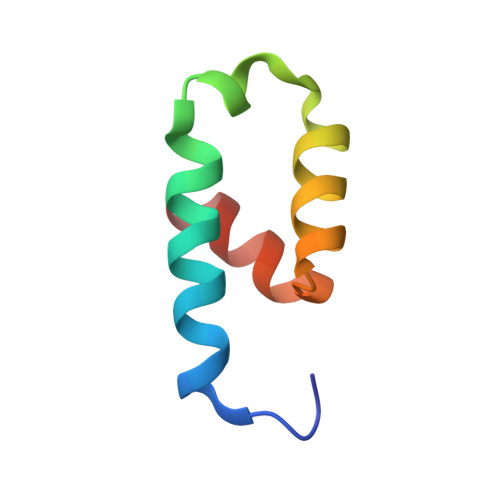
Crystal structure of the conserved N-terminal domain of the peroxisomal matrix protein import receptor, Pex14p
Su, J.R., Takeda, K., Tamura, S., Fujiki, Y., Miki, K.(2009) Proc Natl Acad Sci U S A 106: 417-421
- PubMed: 19122147 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0808681106
- Primary Citation Related Structures:
3FF5 - PubMed Abstract:
Pex14p is a central component of the peroxisomal protein import machinery, in which the conserved N-terminal domain mediates dynamic interactions with other peroxins including Pex5p, Pex13p, and Pex19p. Here, we report the crystal structure of the conserved N-terminal domain of Pex14p with a three-helix bundle. A hydrophobic surface is composed of the conserved residues, of which two phenylalanine residues (Phe-35 and Phe-52) protrude to the solvent. Consequently, two putative binding pockets suitable for recognizing the helical WXXXF/Y motif of Pex5p are formed on the surface by the two phenylalanine residues accompanying with positively charged residues. The structural feature agrees well with our earlier findings where F35A/L36A and F52A/L53A mutants were impaired in the interactions with other peroxins such as Pex5p and Pex13p. Pex14p variants each with Phe-to-Ala mutation at positions 35, 52, and 35/52, respectively, were defective in restoring the impaired peroxisomal protein import in pex14 Chinese hamster ovary mutant ZP161 cells. Moreover, in GST pull-down assays His(6)-Pex5pL bound only to GST-Pex14p(25-70), not to any of GST-Pex14p(25-70)F35A, GST-Pex14p(25-70)F52A, and GST-Pex14p(25-70)F35A/F52A. Endogenous Pex5p was recruited to FLAG-Pex14p on peroxisomes in vivo but barely to FLAG-Pex14pF35A, FLAG-Pex14pF52A, and FLAG-Pex14pF35A/F52A. Collectively, Phe-35 and Phe-52 are essential for the Pex14p functions, including the interaction between Pex14p and Pex5p.
- Department of Chemistry, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto 606-8502, Japan.
Organizational Affiliation: