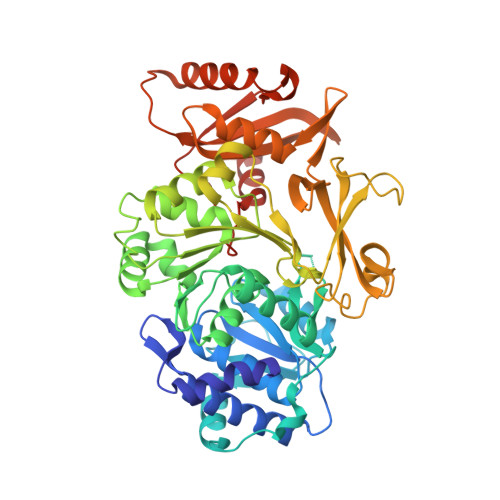
Crystal structure of Bacillus cereus D-alanyl carrier protein ligase (DltA) in complex with ATP.
Osman, K.T., Du, L., He, Y., Luo, Y.(2009) J Mol Biology 388: 345-355
- PubMed: 19324056 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.03.040
- Primary Citation Related Structures:
3FCC, 3FCE - PubMed Abstract:
D-alanylation of lipoteichoic acids modulates the surface charge and ligand binding of the Gram-positive cell wall. Disruption of the bacterial dlt operon involved in teichoic acid alanylation, as well as inhibition of the DltA (D-alanyl carrier protein ligase) protein, has been shown to render the bacterium more susceptible to conventional antibiotics and host defense responses. The DltA catalyzes the adenylation and thiolation reactions of d-alanine. This enzyme belongs to a superfamily of AMP-forming domains such as the ubiquitous acetyl-coenzyme A synthetase. We have determined the 1.9-A-resolution crystal structure of a DltA protein from Bacillus cereus in complex with ATP. This structure sheds light on the geometry of the bound ATP. The invariant catalytic residue Lys492 appears to be mobile, suggesting a molecular mechanism of catalysis for this superfamily of enzymes. Specific roles are also revealed for two other invariant residues: the divalent cation-stabilizing Glu298 and the beta-phosphate-interacting Arg397. Mutant proteins with a glutamine substitution at position 298 or 397 are inactive.
- Department of Biochemistry, University of Saskatchewan, A3 Health Sciences Building, 107 Wiggins Road, Saskatoon, Saskatchewan, Canada S7N 5E5.
Organizational Affiliation: