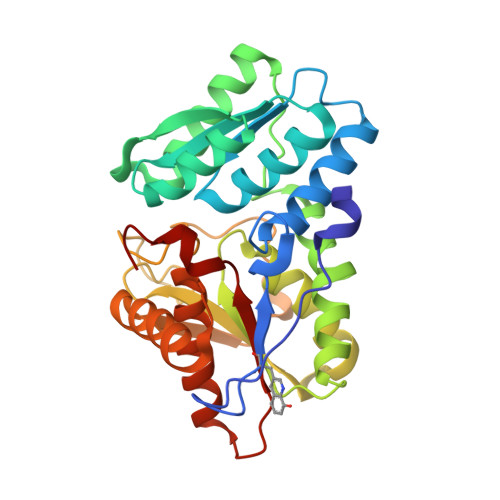
Genetic incorporation of a metal-ion chelating amino acid into proteins as a biophysical probe.
Lee, H.S., Spraggon, G., Schultz, P.G., Wang, F.(2009) J Am Chem Soc 131: 2481-2483
- PubMed: 19193005 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja808340b
- Primary Citation Related Structures:
3FCA - PubMed Abstract:
A metal-ion chelating amino acid, (8-hydroxyquinolin-3-yl)alanine, was genetically encoded in E. coli by an amber nonsense codon and corresponding orthogonal tRNA/aminoacyl-tRNA synthetase pair. The amino acid was incorporated into TM0665 protein, and the mutant protein was cocrystallized with Zn(2+) to determine the structure by SAD phasing. The structure showed a high occupancy of the heavy metal bound to the HQ-Ala residue, and the heavy metal provided excellent phasing power to determine the structure. This method also facilitates the de novo design of metalloproteins with novel structures and functions, including fluorescent sensors.
- Department of Chemistry, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, California 92037, USA.
Organizational Affiliation: