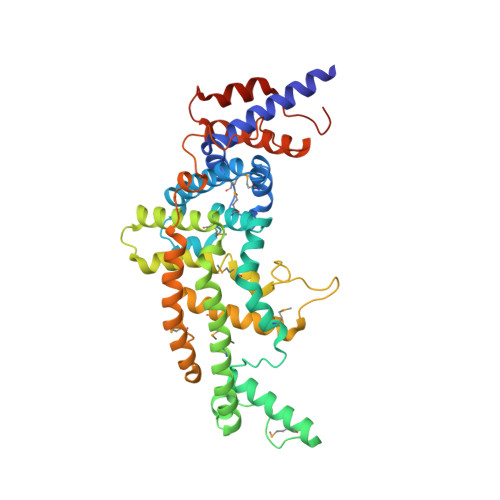
Crystal structure of the GTPase-activating protein-related domain from IQGAP1.
Kurella, V.B., Richard, J.M., Parke, C.L., Lecour, L.F., Bellamy, H.D., Worthylake, D.K.(2009) J Biological Chem 284: 14857-14865
- PubMed: 19321438 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M808974200
- Primary Citation Related Structures:
3FAY - PubMed Abstract:
IQGAP1 is a 190-kDa molecular scaffold containing several domains required for interaction with numerous proteins. One domain is homologous to Ras GTPase-activating protein (GAP) domains. However, instead of accelerating hydrolysis of bound GTP on Ras IQGAP1, using its GAP-related domain (GRD) binds to Cdc42 and Rac1 and stabilizes their GTP-bound states. We report here the crystal structure of the isolated IQGAP1 GRD. Despite low sequence conservation, the overall structure of the GRD is very similar to the GAP domains from p120 RasGAP, neurofibromin, and SynGAP. However, instead of the catalytic "arginine finger" seen in functional Ras GAPs, the GRD has a conserved threonine residue. GRD residues 1099-1129 have no structural equivalent in RasGAP and are seen to form an extension at one end of the molecule. Because the sequence of these residues is highly conserved, this region likely confers a functionality particular to IQGAP family GRDs. We have used isothermal titration calorimetry to demonstrate that the isolated GRD binds to active Cdc42. Assuming a mode of interaction similar to that displayed in the Ras-RasGAP complex, we created an energy-minimized model of Cdc42.GTP bound to the GRD. Residues of the GRD that contact Cdc42 map to the surface of the GRD that displays the highest level of sequence conservation. The model indicates that steric clash between threonine 1046 with the phosphate-binding loop and other subtle changes would likely disrupt the proper geometry required for GTP hydrolysis.
- Department of Biochemistry and Molecular Biology, Louisiana State University Health Sciences Center, New Orleans, Louisiana 70112, USA.
Organizational Affiliation: