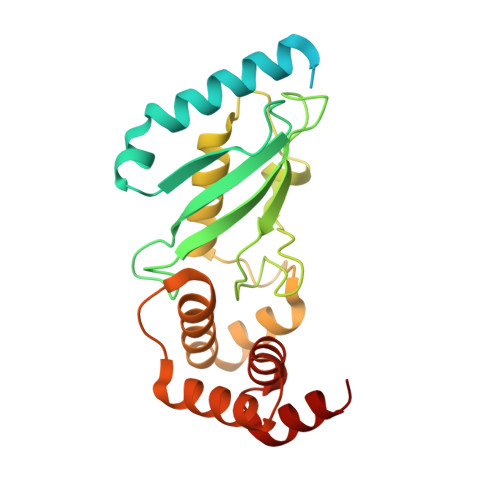
Structure of full-length ubiquitin-conjugating enzyme E2-25K (huntingtin-interacting protein 2).
Wilson, R.C., Hughes, R.C., Flatt, J.W., Meehan, E.J., Ng, J.D., Twigg, P.D.(2009) Acta Crystallogr Sect F Struct Biol Cryst Commun 65: 440-444
- PubMed: 19407372 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309109011117
- Primary Citation Related Structures:
3E46, 3F92 - PubMed Abstract:
The ubiquitin-conjugating enzyme E2-25K has been identified as a huntingtin (the key protein in Huntington's disease) interacting protein and has been shown to play a role in mediating the toxicity of Abeta, the principal protein involved in Alzheimer's disease pathogenesis. E2-25K is a dual-domain protein with an ubiquitin-associated (UBA) domain as well as a conserved ubiquitin-conjugating (UBC) domain which catalyzes the formation of a covalent bond between the C-terminal glycine of an ubiquitin molecule and the -amine of a lysine residue on the acceptor protein as part of the ubiquitin-proteasome pathway. The crystal structures of E2-25K M172A mutant protein at pH 6.5 and pH 8.5 were determined to 1.9 and 2.2 A resolution, respectively. Examination of the structures revealed domain-domain interactions between the UBC and UBA domains which have not previously been reported.
- University of Alabama in Huntsville, 35899, USA.
Organizational Affiliation: