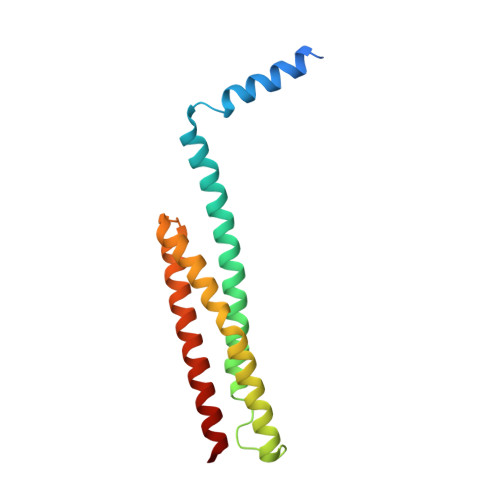
Crystal structure of the nonerythroid alpha-spectrin tetramerization site reveals differences between erythroid and nonerythroid spectrin tetramer formation.
Mehboob, S., Song, Y., Witek, M., Long, F., Santarsiero, B.D., Johnson, M.E., Fung, L.W.(2010) J Biological Chem 285: 14572-14584
- PubMed: 20228407 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.080028
- Primary Citation Related Structures:
3F31 - PubMed Abstract:
We have solved the crystal structure of a segment of nonerythroid alpha-spectrin (alphaII) consisting of the first 147 residues to a resolution of 2.3 A. We find that the structure of this segment is generally similar to a corresponding segment from erythroid alpha-spectrin (alphaI) but exhibits unique differences with functional significance. Specific features include the following: (i) an irregular and frayed first helix (Helix C'); (ii) a helical conformation in the junction region connecting Helix C' with the first structural domain (D1); (iii) a long A(1)B(1) loop in D1; and (iv) specific inter-helix hydrogen bonds/salt bridges that stabilize D1. Our findings suggest that the hydrogen bond networks contribute to structural domain stability, and thus rigidity, in alphaII, and the lack of such hydrogen bond networks in alphaI leads to flexibility in alphaI. We have previously shown the junction region connecting Helix C' to D1 to be unstructured in alphaI (Park, S., Caffrey, M. S., Johnson, M. E., and Fung, L. W. (2003) J. Biol. Chem. 278, 21837-21844) and now find it to be helical in alphaII, an important difference for alpha-spectrin association with beta-spectrin in forming tetramers. Homology modeling and molecular dynamics simulation studies of the structure of the tetramerization site, a triple helical bundle of partial domain helices, show that mutations in alpha-spectrin will affect Helix C' structural flexibility and/or the junction region conformation and may alter the equilibrium between spectrin dimers and tetramers in cells. Mutations leading to reduced levels of functional tetramers in cells may potentially lead to abnormal neuronal functions.
- Center for Pharmaceutical Biotechnology, University of Illinois at Chicago, Chicago, Illinois 60607, USA. shahila@uic.edu
Organizational Affiliation: