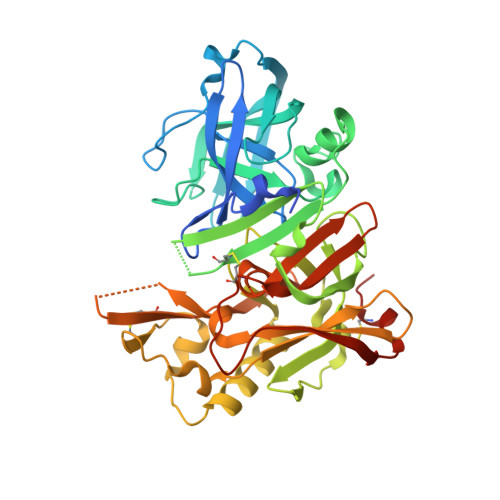
Identification of a small molecule beta-secretase inhibitor that binds without catalytic aspartate engagement.
Steele, T.G., Hills, I.D., Nomland, A.A., de Leon, P., Allison, T., McGaughey, G., Colussi, D., Tugusheva, K., Haugabook, S.J., Espeseth, A.S., Zuck, P., Graham, S.L., Stachel, S.J.(2009) Bioorg Med Chem Lett 19: 17-20
- PubMed: 19036583
- DOI: https://doi.org/10.1016/j.bmcl.2008.11.027
- Primary Citation Related Structures:
3EXO - PubMed Abstract:
A small molecule inhibitor of beta-secretase with a unique binding mode has been developed. Crystallographic determination of the enzyme-inhibitor complex shows the catalytic aspartate residues in the active site are not engaged in inhibitor binding. This unprecedented binding mode in the field of aspartyl protease inhibition is described.
- Department of Medicinal Chemistry, Merck Research Laboratories, Merck & Co, West Point, PA 19486, USA.
Organizational Affiliation: