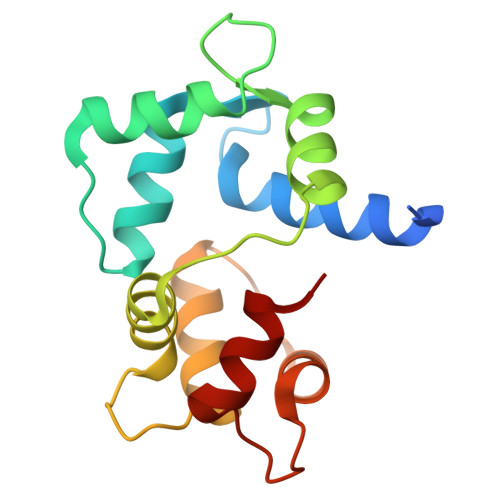
Structural insights into the mechanism of calmodulin binding to death receptors.
Cao, P., Zhang, W., Gui, W., Dong, Y., Jiang, T., Gong, Y.(2014) Acta Crystallogr D Biol Crystallogr 70: 1604-1613
- PubMed: 24914971 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004714006919
- Primary Citation Related Structures:
3EWT, 3EWV - PubMed Abstract:
The death receptors Fas, p75(NTR) and DR6 are key components of extrinsically activated apoptosis. Characterization of how they interact with the adaptors is crucial in order to unravel the signalling mechanisms. However, the exact conformation that their intracellular death domain adopts upon binding downstream partners remains unclear. One model suggests that it adopts a typical compact fold, whilst a second model proposed an open conformation. Calmodulin (CaM), a major calcium sensor, has previously been reported to be one of the Fas adaptors that modulate apoptosis. This work reports that CaM also binds directly to the death domains of p75(NTR) and DR6, indicating that it serves as a common modulator of the death receptors. Two crystal structures of CaM in complexes with the corresponding binding regions of Fas and p75(NTR) are also reported. Interestingly, the precise CaM-binding sites were mapped to different regions: helix 1 in Fas and helix 5 in p75(NTR) and DR6. A novel 1-11 motif for CaM binding was observed in p75(NTR). Modelling the complexes of CaM with full-length receptors reveals that the opening of the death domains would be essential in order to expose their binding sites for CaM. These results may facilitate understanding of the diverse functional repertoire of death receptors and CaM and provide further insights necessary for the design of potential therapeutic peptide agents.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Chaoyang District, Beijing 100101, People's Republic of China.
Organizational Affiliation: