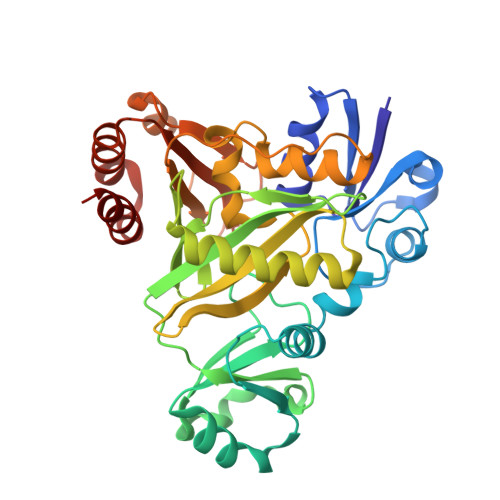
Structural analysis of the active site geometry of N(5)-Carboxyaminoimidazole ribonucleotide synthetase from Escherichia coli.
Thoden, J.B., Holden, H.M., Firestine, S.M.(2008) Biochemistry 47: 13346-13353
- PubMed: 19053251 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi801734z
- Primary Citation Related Structures:
3ETH, 3ETJ - PubMed Abstract:
N(5)-Carboxyaminoimidazole ribonucleotide synthetase (N(5)-CAIR synthetase) converts 5-aminoimidazole ribonucleotide (AIR), MgATP, and bicarbonate into N(5)-CAIR, MgADP, and P(i). The enzyme is required for de novo purine biosynthesis in microbes yet is not found in humans suggesting that it represents an ideal and unexplored target for antimicrobial drug design. Here we report the X-ray structures of N(5)-CAIR synthetase from Escherichia coli with either MgATP or MgADP/P(i) bound in the active site cleft. These structures, determined to 1.6-A resolution, provide detailed information regarding the active site geometry before and after ATP hydrolysis. In both structures, two magnesium ions are observed. Each of these is octahedrally coordinated, and the carboxylate side chain of Glu238 bridges them. For the structure of the MgADP/P(i) complex, crystals were grown in the presence of AIR and MgATP. No electron density was observed for AIR, and the electron density corresponding to the nucleotide clearly revealed the presence of ADP and P(i) rather than ATP. The bound P(i) shifts by approximately 3 A relative to the gamma-phosphoryl group of ATP and forms electrostatic interactions with the side chains of Arg242 and His244. Since the reaction mechanism of N(5)-CAIR synthetase is believed to proceed via a carboxyphosphate intermediate, we propose that the location of the inorganic phosphate represents the binding site for stabilization of this reactive species. Using the information derived from the two structures reported here, coupled with molecular modeling, we propose a catalytic mechanism for N(5)-CAIR synthetase.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.
Organizational Affiliation: