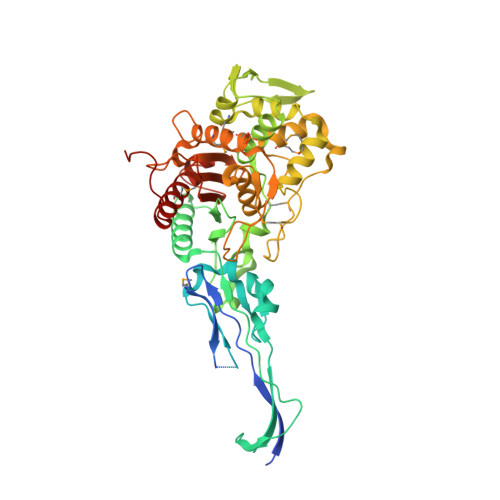
Crystal Structures of Penicillin-binding Protein 2 from Penicillin-susceptible and -resistant Strains of Neisseria gonorrhoeae Reveal an Unexpectedly Subtle Mechanism for Antibiotic Resistance.
Powell, A.J., Tomberg, J., Deacon, A.M., Nicholas, R.A., Davies, C.(2009) J Biological Chem 284: 1202-1212
- PubMed: 18986991 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M805761200
- Primary Citation Related Structures:
3EQU, 3EQV - PubMed Abstract:
Penicillin-binding protein 2 (PBP2) from N. gonorrhoeae is the major molecular target for beta-lactam antibiotics used to treat gonococcal infections. PBP2 from penicillin-resistant strains of N. gonorrhoeae harbors an aspartate insertion after position 345 (Asp-345a) and 4-8 additional mutations, but how these alter the architecture of the protein is unknown. We have determined the crystal structure of PBP2 derived from the penicillin-susceptible strain FA19, which shows that the likely effect of Asp-345a is to alter a hydrogen-bonding network involving Asp-346 and the SXN triad at the active site. We have also solved the crystal structure of PBP2 derived from the penicillin-resistant strain FA6140 that contains four mutations near the C terminus of the protein. Although these mutations lower the second order rate of acylation for penicillin by 5-fold relative to wild type, comparison of the two structures shows only minor structural differences, with the positions of the conserved residues in the active site essentially the same in both. Kinetic analyses indicate that two mutations, P551S and F504L, are mainly responsible for the decrease in acylation rate. Melting curves show that the four mutations lower the thermal stability of the enzyme. Overall, these data suggest that the molecular mechanism underlying antibiotic resistance contributed by the four mutations is subtle and involves a small but measurable disordering of residues in the active site region that either restricts the binding of antibiotic or impedes conformational changes that are required for acylation by beta-lactam antibiotics.
- Department of Biochemistry and Molecular Biology, Medical University of South Carolina, Charleston, South Carolina 29425, USA.
Organizational Affiliation: