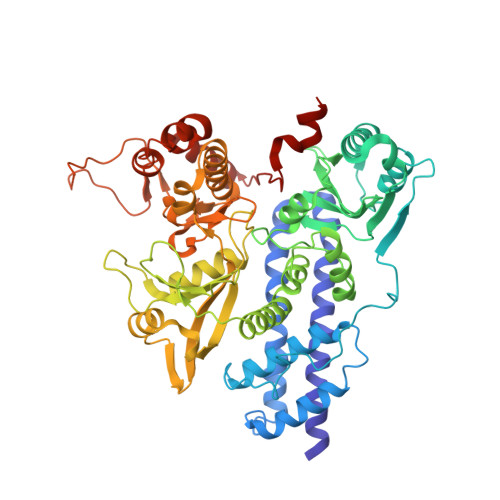
Structure of the bifunctional isocitrate dehydrogenase kinase/phosphatase.
Zheng, J., Jia, Z.(2010) Nature 465: 961-965
- PubMed: 20505668 Search on PubMed
- DOI: https://doi.org/10.1038/nature09088
- Primary Citation Related Structures:
3EPS, 3LC6, 3LCB - PubMed Abstract:
The Escherichia coli isocitrate dehydrogenase kinase/phosphatase (AceK) is a unique bifunctional enzyme that phosphorylates or dephosphorylates isocitrate dehydrogenase (ICDH) in response to environmental changes, resulting in the inactivation or, respectively, activation of ICDH. ICDH inactivation short-circuits the Krebs cycle by enabling the glyoxlate bypass. It was the discovery of AceK and ICDH that established the existence of protein phosphorylation regulation in prokaryotes. As a 65-kDa protein, AceK is significantly larger than typical eukaryotic protein kinases. Apart from the ATP-binding motif, AceK does not share sequence homology with any eukaryotic protein kinase or phosphatase. Most intriguingly, AceK possesses the two opposing activities of protein kinase and phosphatase within one protein, and specifically recognizes only intact ICDH. Additionally, AceK has strong ATPase activity. It has been shown that AceK kinase, phosphatase and ATPase activities reside at the same site, although the molecular basis of such multifunctionality and its regulation remains completely unknown. Here we report the structures of AceK and its complex with ICDH. The AceK structure reveals a eukaryotic protein-kinase-like domain containing ATP and a regulatory domain with a novel fold. As an AceK phosphatase activator and kinase inhibitor, AMP is found to bind in an allosteric site between the two AceK domains. An AMP-mediated conformational change exposes and shields ATP, acting as a switch between AceK kinase and phosphatase activities, and ICDH-binding induces further conformational change for AceK activation. The substrate recognition loop of AceK binds to the ICDH dimer, allowing higher-order substrate recognition and interaction, and inducing critical conformational change at the phosphorylation site of ICDH.
- Department of Biochemistry, Queen's University, Kingston, Ontario K7L 3N6, Canada.
Organizational Affiliation: