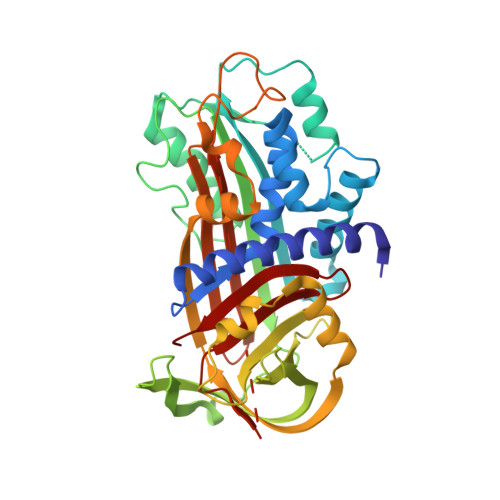
High quality structure of cleaved PAI-1-stab.
Dewilde, M., Strelkov, S.V., Rabijns, A., Declerck, P.J.(2009) J Struct Biol 165: 126-132
- PubMed: 19059484 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2008.11.001
- Primary Citation Related Structures:
3EOX - PubMed Abstract:
Here we report the crystal structure of a stablilized plasminogen activator inhibitor-1 variant (PAI-1-N150H-K154T-Q301P-Q319L-M354I (PAI-1-stab)) that shows a cleavage within the reactive centre loop. The new structure is of superior quality compared to the previously determined structure of the cleaved PAI-1-A335P mutant. We present a detailed comparison of the two structures and also compare them with the structure of the active PAI-1-stab. The structural data give important insights into the working mechanism of PAI-1 and also explain the role of various stabilizing mutations.
- Katholieke Universiteit Leuven, Belgium.
Organizational Affiliation: