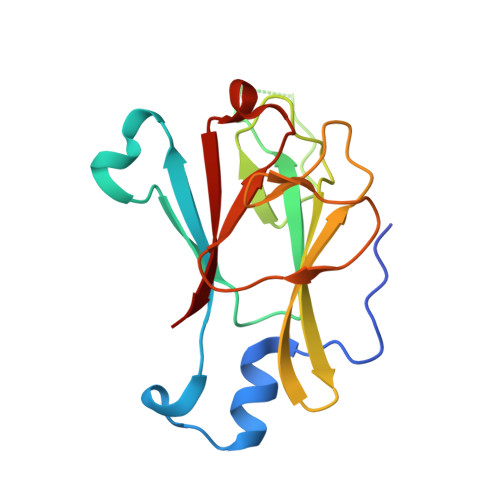
Crystal structure of the Pml1p subunit of the yeast precursor mRNA retention and splicing complex.
Trowitzsch, S., Weber, G., Luhrmann, R., Wahl, M.C.(2009) J Mol Biology 385: 531-541
- PubMed: 19010333 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.10.087
- Primary Citation Related Structures:
3ELS, 3ELV - PubMed Abstract:
The precursor mRNA retention and splicing (RES) complex mediates nuclear retention and enhances splicing of precursor mRNAs. The RES complex from yeast comprises three proteins, Snu17p, Bud13p and Pml1p. Snu17p acts as a central platform that concomitantly binds the Bud13p and Pml1p subunits via short peptide epitopes. As a step to decipher the molecular architecture of the RES complex, we have determined crystal structures of full-length Pml1p and N-terminally truncated Pml1p. The first 50 residues of full-length Pml1p, encompassing the Snu17p-binding region, are disordered, showing that Pml1p binds to Snu17p via an intrinsically unstructured region. The remainder of Pml1p folds as a forkhead-associated (FHA) domain, which is expanded by a number of noncanonical elements compared with known FHA domains from other proteins. An atypical N-terminal appendix runs across one beta-sheet and thereby stabilizes the domain as shown by deletion experiments. FHA domains are thought to constitute phosphopeptide-binding elements. Consistently, a sulfate ion was found at the putative phosphopeptide-binding loops of full-length Pml1p. The N-terminally truncated version of the protein lacked a similar phosphopeptide mimic but retained an almost identical structure. A long loop neighboring the putative phosphopeptide-binding site was disordered in both structures. Comparison with other FHA domain proteins suggests that this loop adopts a defined conformation upon ligand binding and thereby confers ligand specificity. Our results show that in the RES complex, an FHA domain of Pml1p is flexibly tethered via an unstructured N-terminal region to Snu17p.
- Zelluläre Biochemie, Max-Planck-Institut für Biophysikalische Chemie, Am Fassberg 11, D-37077 Göttingen, Germany.
Organizational Affiliation: