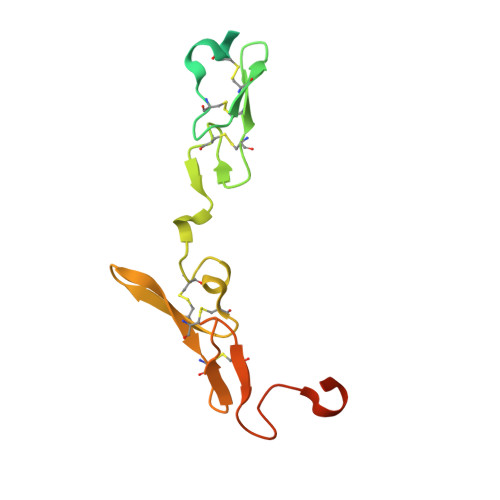
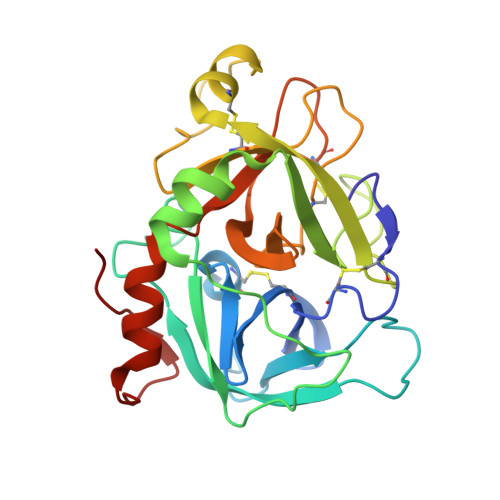
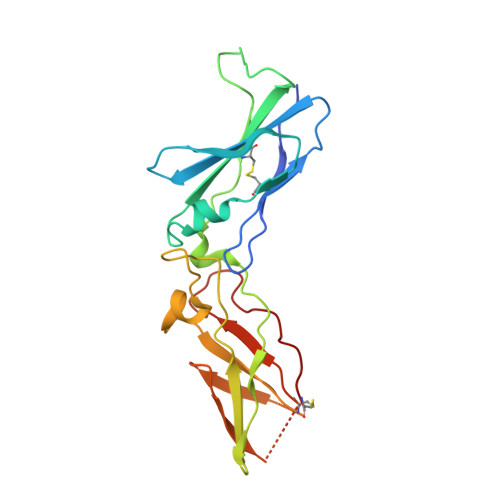
Mechanism of the Ca2+-induced enhancement of the intrinsic factor VIIa activity
Bjelke, J.R., Olsen, O.H., Fodje, M., Svensson, L.A., Bang, S., Bolt, G., Kragelund, B.B., Persson, E.(2008) J Biological Chem 283: 25863-25870
- PubMed: 18640965 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M800841200
- Primary Citation Related Structures:
3ELA - PubMed Abstract:
The intrinsic activity of coagulation factor VIIa (FVIIa) is dependent on Ca(2+) binding to a loop (residues 210-220) in the protease domain. Structural analysis revealed that Ca(2+) may enhance the activity by attenuating electrostatic repulsion of Glu(296) and/or by facilitating interactions between the loop and Lys(161) in the N-terminal tail. In support of the first mechanism, the mutations E296V and D212N resulted in similar, about 2-fold, enhancements of the amidolytic activity. Moreover, mutation of the Lys(161)-interactive residue Asp(217) or Asp(219) to Ala reduced the amidolytic activity by 40-50%, whereas the K161A mutation resulted in 80% reduction. Hence one of these Asp residues in the Ca(2+)-binding loop appears to suffice for some residual interaction with Lys(161), whereas the more severe effect upon replacement of Lys(161) is due to abrogation of the interaction with the N-terminal tail. However, Ca(2+) attenuation of the repulsion between Asp(212) and Glu(296) keeps the activity above that of apoFVIIa. Altogether, our data suggest that repulsion involving Asp(212) in the Ca(2+)-binding loop suppresses FVIIa activity and that optimal activity requires a favorable interaction between the Ca(2+)-binding loop and the N-terminal tail. Crystal structures of tissue factor-bound FVIIa(D212N) and FVIIa(V158D/E296V/M298Q) revealed altered hydrogen bond networks, resembling those in factor Xa and thrombin, after introduction of the D212N and E296V mutations plausibly responsible for tethering the N-terminal tail to the activation domain. The charge repulsion between the Ca(2+)-binding loop and the activation domain appeared to be either relieved by charge removal and new hydrogen bonds (D212N) or abolished (E296V). We propose that Ca(2+) stimulates the intrinsic FVIIa activity by a combination of charge neutralization and loop stabilization.
- Department of Protein Structure and Biophysics, Novo Nordisk A/S, Novo Nordisk Park, DK-2760 Måløv, Denmark.
Organizational Affiliation: