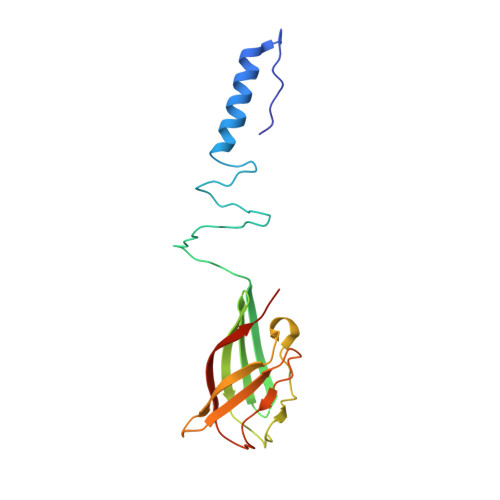
Structure and molecular assignment of lactococcal phage TP901-1 baseplate.
Bebeacua, C., Bron, P., Lai, L., Vegge, C.S., Brondsted, L., Spinelli, S., Campanacci, V., Veesler, D., van Heel, M., Cambillau, C.(2010) J Biological Chem 285: 39079-39086
- PubMed: 20937834 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.175646
- Primary Citation Related Structures:
3EJC - PubMed Abstract:
P335 lactococcal phages infect the gram(+) bacterium Lactococcus lactis using a large multiprotein complex located at the distal part of the tail and termed baseplate (BP). The BP harbors the receptor-binding proteins (RBPs), which allow the specific recognition of saccharidic receptors localized on the host cell surface. We report here the electron microscopic structure of the phage TP901-1 wild-type BP as well as those of two mutants bppL (-) and bppU(-), lacking BppL (the RBPs) or both peripheral BP components (BppL and BppU), respectively. We also achieved an electron microscopic reconstruction of a partial BP complex, formed by BppU and BppL. This complex exhibits a tripod shape and is composed of nine BppLs and three BppUs. These structures, combined with light-scattering measurements, led us to propose that the TP901-1 BP harbors six tripods at its periphery, located around the central tube formed by ORF46 (Dit) hexamers, at its proximal end, and a ORF47 (Tal) trimer at its distal extremity. A total of 54 BppLs (18 RBPs) are thus available to mediate host anchoring with a large apparent avidity. TP901-1 BP exhibits an infection-ready conformation and differs strikingly from the lactococcal phage p2 BP, bearing only 6 RBPs, and which needs a conformational change to reach its activated state. The comparison of several Siphoviridae structures uncovers a close organization of their central BP core whereas striking differences occur at the periphery, leading to diverse mechanisms of host recognition.
- Department of Biological Sciences, Imperial College London, South Kensington Campus, London SW7 2AZ, United Kingdom.
Organizational Affiliation: