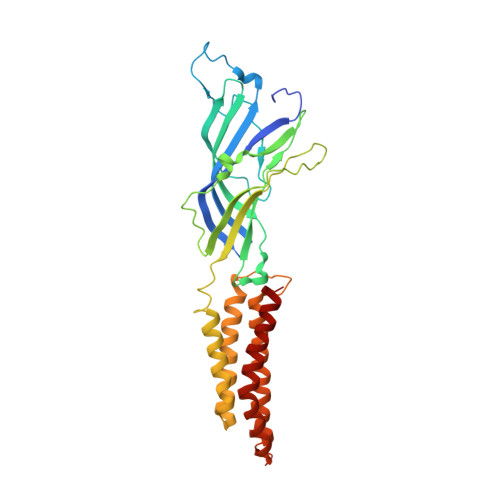
Structure of a potentially open state of a proton-activated pentameric ligand-gated ion channel
Hilf, R.J., Dutzler, R.(2008) Nature 457: 115-118
- PubMed: 18987630 Search on PubMed
- DOI: https://doi.org/10.1038/nature07461
- Primary Citation Related Structures:
3EHZ, 3EI0 - PubMed Abstract:
The X-ray structure of a pentameric ligand-gated ion channel from Erwinia chrysanthemi (ELIC) has recently provided structural insight into this family of ion channels at high resolution. The structure shows a homo-pentameric protein with a barrel-stave architecture that defines an ion-conduction pore located on the fivefold axis of symmetry. In this structure, the wide aqueous vestibule that is encircled by the extracellular ligand-binding domains of the five subunits narrows to a discontinuous pore that spans the lipid bilayer. The pore is constricted by bulky hydrophobic residues towards the extracellular side, which probably serve as barriers that prevent the diffusion of ions. This interrupted pore architecture in ELIC thus depicts a non-conducting conformation of a pentameric ligand-gated ion channel, the thermodynamically stable state in the absence of bound ligand. As ligand binding promotes pore opening in these ion channels and the specific ligand for ELIC has not yet been identified, we have turned our attention towards a homologous protein from the cyanobacterium Gloebacter violaceus (GLIC). GLIC was shown to form proton-gated channels that are activated by a pH decrease on the extracellular side and that do not desensitize after activation. Both prokaryotic proteins, ELIC and GLIC form ion channels that are selective for cations over anions with poor discrimination among monovalent cations, characteristics that resemble the conduction properties of the cation-selective branch of the family that includes acetylcholine and serotonin receptors. Here we present the X-ray structure of GLIC at 3.1 A resolution. The structure reveals a conformation of the channel that is distinct from ELIC and that probably resembles the open state. In combination, both structures suggest a novel gating mechanism for pentameric ligand-gated ion channels where channel opening proceeds by a change in the tilt of the pore-forming helices.
- Department of Biochemistry, University of Zürich, Winterthurerstrasse 190, CH-8057 Zürich, Switzerland.
Organizational Affiliation: