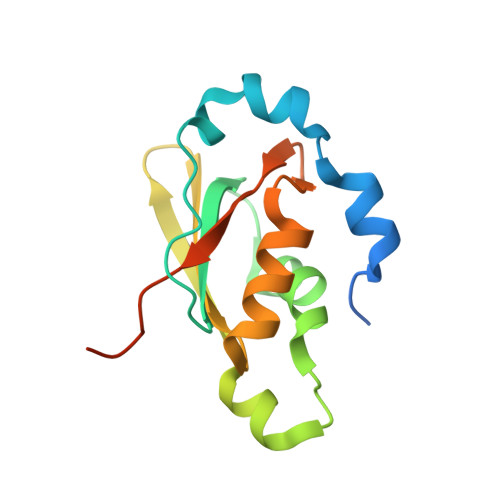
Functional stabilization of an RNA recognition motif by a noncanonical N-terminal expansion
Netter, C., Weber, G., Benecke, H., Wahl, M.C.(2009) RNA 15: 1305-1313
- PubMed: 19447915 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.1359909
- Primary Citation Related Structures:
3EGN - PubMed Abstract:
RNA recognition motifs (RRMs) constitute versatile macromolecular interaction platforms. They are found in many components of spliceosomes, in which they mediate RNA and protein interactions by diverse molecular strategies. The human U11/U12-65K protein of the minor spliceosome employs a C-terminal RRM to bind hairpin III of the U12 small nuclear RNA (snRNA). This interaction comprises one side of a molecular bridge between the U11 and U12 small nuclear ribonucleoprotein particles (snRNPs) and is reminiscent of the binding of the N-terminal RRMs in the major spliceosomal U1A and U2B'' proteins to hairpins in their cognate snRNAs. Here we show by mutagenesis and electrophoretic mobility shift assays that the beta-sheet surface and a neighboring loop of 65K C-terminal RRM are involved in RNA binding, as previously seen in canonical RRMs like the N-terminal RRMs of the U1A and U2B'' proteins. However, unlike U1A and U2B'', some 30 residues N-terminal of the 65K C-terminal RRM core are additionally required for stable U12 snRNA binding. The crystal structure of the expanded 65K C-terminal RRM revealed that the N-terminal tail adopts an alpha-helical conformation and wraps around the protein toward the face opposite the RNA-binding platform. Point mutations in this part of the protein had only minor effects on RNA affinity. Removal of the N-terminal extension significantly decreased the thermal stability of the 65K C-terminal RRM. These results demonstrate that the 65K C-terminal RRM is augmented by an N-terminal element that confers stability to the domain, and thereby facilitates stable RNA binding.
- Max-Planck-Institut für Biophysikalische Chemie, D-37077 Göttingen, Germany.
Organizational Affiliation: