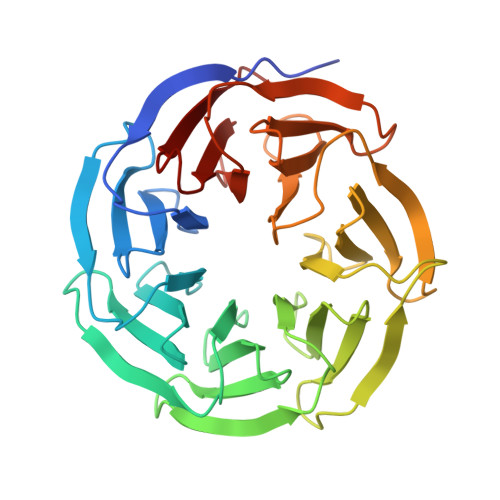
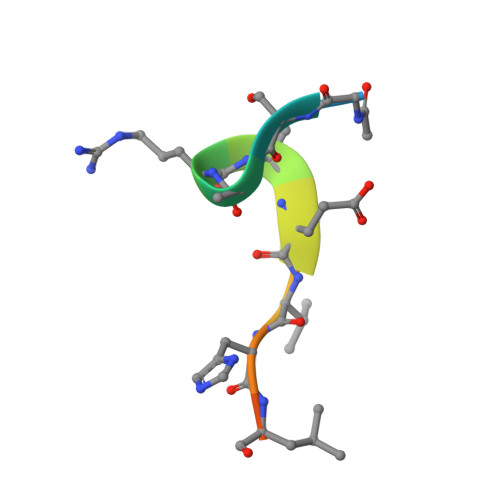
Structure of WDR5 bound to mixed lineage leukemia protein-1 peptide.
Patel, A., Dharmarajan, V., Cosgrove, M.S.(2008) J Biological Chem 283: 32158-32161
- PubMed: 18829459 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.C800164200
- Primary Citation Related Structures:
3EG6 - PubMed Abstract:
The mixed lineage leukemia protein-1 (MLL1) catalyzes histone H3 lysine 4 methylation and is regulated by interaction with WDR5 (WD-repeat protein-5), RbBP5 (retinoblastoma-binding protein-5), and the Ash2L (absent, small, homeotic discs-2-like) oncoprotein. In the accompanying investigation, we describe the identification of a conserved arginine containing motif, called the "Win" or WDR5 interaction motif, that is essential for the assembly and H3K4 dimethylation activity of the MLL1 core complex. Here we present a 1.7-A crystal structure of WDR5 bound to a peptide derived from the MLL1 Win motif. Our results show that Arg-3765 of MLL1 is bound in the same arginine binding pocket on WDR5 that was previously suggested to bind histone H3. Thermodynamic binding experiments show that the MLL1 Win peptide is preferentially recognized by WDR5. These results are consistent with a model in which WDR5 recognizes Arg-3765 of MLL1, which is essential for the assembly and enzymatic activity of the MLL1 core complex.
- Department of Biology, Syracuse University, Syracuse, New York 13244, USA.
Organizational Affiliation: