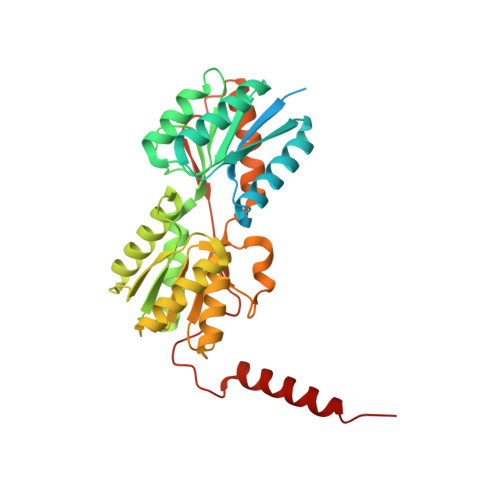
Crystal structure of a 1.6-hexanediol bound tetrameric form of Escherichia coli lac-repressor refined to 2.1 A resolution
Stenberg, K.A.E., Vihinen, M.(2009) Proteins 75: 748-759
- PubMed: 19004002 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22284
- Primary Citation Related Structures:
3EDC - PubMed Abstract:
We report the structure of a novel tetrameric form of the lactose repressor (LacI) protein from Escherichia coli refined to 2.1 A resolution. The tetramer is bound to 1.6-hexanediol present in the crystallization solution and the final R(free) for the structure is 0.201. The structure confirms previously reported structures on the monomer level. However, the tetramer is much more densely packed. This adds a new level of complexity to the interpretation of mutational effects and challenges details in the current model for LacI function. Several amino acids, previously associated with changes in function but unexplained at the structural level, appear in a new structural context in this tetramer which provides new implications for their function.
- Faculty of Biosciences, Division of Biochemistry, University of Helsinki, Helsinki, Finland. kaj.stenberg@helsinki.fi
Organizational Affiliation: